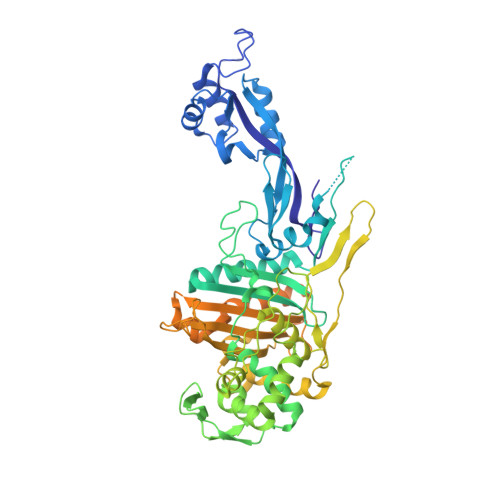
Integrative structural biology of the penicillin-binding protein-1 from Staphylococcus aureus , an essential component of the divisome machinery.
Martinez-Caballero, S., Mahasenan, K.V., Kim, C., Molina, R., Feltzer, R., Lee, M., Bouley, R., Hesek, D., Fisher, J.F., Munoz, I.G., Chang, M., Mobashery, S., Hermoso, J.A.(2021) Comput Struct Biotechnol J 19: 5392-5405
- PubMed: 34667534 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.csbj.2021.09.018
- Primary Citation Related Structures:
7O49, 7O4A, 7O4B, 7O4C, 7OK9 - PubMed Abstract:
The penicillin-binding proteins are the enzyme catalysts of the critical transpeptidation crosslinking polymerization reaction of bacterial peptidoglycan synthesis and the molecular targets of the penicillin antibiotics. Here, we report a combined crystallographic, small-angle X-ray scattering (SAXS) in-solution structure, computational and biophysical analysis of PBP1 of Staphylococcus aureus (sa PBP1), providing mechanistic clues about its function and regulation during cell division. The structure reveals the pedestal domain, the transpeptidase domain, and most of the linker connecting to the "penicillin-binding protein and serine/threonine kinase associated" (PASTA) domains, but not its two PASTA domains, despite their presence in the construct. To address this absence, the structure of the PASTA domains was determined at 1.5 Å resolution. Extensive molecular-dynamics simulations interpret the PASTA domains of sa PBP1 as conformationally mobile and separated from the transpeptidase domain. This conclusion was confirmed by SAXS experiments on the full-length protein in solution. A series of crystallographic complexes with β-lactam antibiotics (as inhibitors) and penta-Gly (as a substrate mimetic) allowed the molecular characterization of both inhibition by antibiotics and binding for the donor and acceptor peptidoglycan strands. Mass-spectrometry experiments with synthetic peptidoglycan fragments revealed binding by PASTA domains in coordination with the remaining domains. The observed mobility of the PASTA domain in sa PBP1 could play a crucial role for in vivo interaction with its glycosyltransferase partner in the membrane or with other components of the divisome machinery, as well as for coordination of transpeptidation and polymerization processes in the bacterial divisome.
- Department of Crystallography and Structural Biology, Institute of Physical Chemistry "Rocasolano", CSIC, 28006 Madrid, Spain.
Organizational Affiliation: