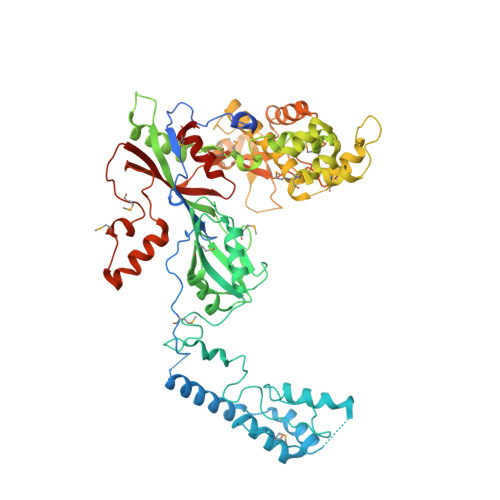
Pseudouridine synthase 7 is an opportunistic enzyme that binds and modifies substrates with diverse sequences and structures.
Purchal, M.K., Eyler, D.E., Tardu, M., Franco, M.K., Korn, M.M., Khan, T., McNassor, R., Giles, R., Lev, K., Sharma, H., Monroe, J., Mallik, L., Koutmos, M., Koutmou, K.S.(2022) Proc Natl Acad Sci U S A 119
- PubMed: 35058356
- DOI: https://doi.org/10.1073/pnas.2109708119
- Primary Citation Related Structures:
7MZV - PubMed Abstract:
Pseudouridine (Ψ) is a ubiquitous RNA modification incorporated by pseudouridine synthase (Pus) enzymes into hundreds of noncoding and protein-coding RNA substrates. Here, we determined the contributions of substrate structure and protein sequence to binding and catalysis by pseudouridine synthase 7 (Pus7), one of the principal messenger RNA (mRNA) modifying enzymes. Pus7 is distinct among the eukaryotic Pus proteins because it modifies a wider variety of substrates and shares limited homology with other Pus family members. We solved the crystal structure of Saccharomyces cerevisiae Pus7, detailing the architecture of the eukaryotic-specific insertions thought to be responsible for the expanded substrate scope of Pus7. Additionally, we identified an insertion domain in the protein that fine-tunes Pus7 activity both in vitro and in cells. These data demonstrate that Pus7 preferentially binds substrates possessing the previously identified UG U AR (R = purine) consensus sequence and that RNA secondary structure is not a strong requirement for Pus7-binding. In contrast, the rate constants and extent of Ψ incorporation are more influenced by RNA structure, with Pus7 modifying UG U AR sequences in less-structured contexts more efficiently both in vitro and in cells. Although less-structured substrates were preferred, Pus7 fully modified every transfer RNA, mRNA, and nonnatural RNA containing the consensus recognition sequence that we tested. Our findings suggest that Pus7 is a promiscuous enzyme and lead us to propose that factors beyond inherent enzyme properties (e.g., enzyme localization, RNA structure, and competition with other RNA-binding proteins) largely dictate Pus7 substrate selection.
- Program in Chemical Biology, University of Michigan, Ann Arbor, MI 48109.
Organizational Affiliation: