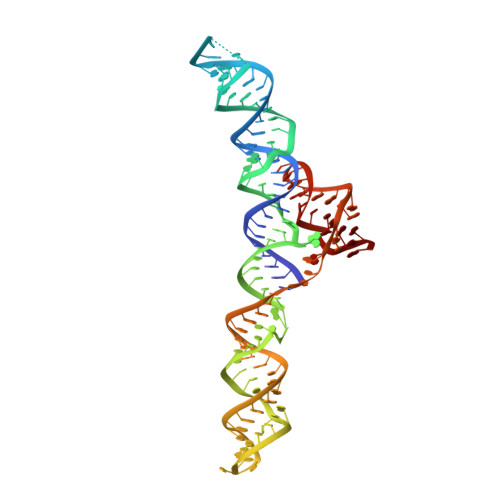
An uncommon [K + (Mg 2+ ) 2 ] metal ion triad imparts stability and selectivity to the Guanidine-I riboswitch.
Trachman 3rd, R.J., Ferre-D'Amare, A.R.(2021) RNA 27: 1257-1264
- PubMed: 34257148 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.078824.121
- Primary Citation Related Structures:
7MLW - PubMed Abstract:
The widespread ykkC -I riboswitch class exemplifies divergent riboswitch evolution. To analyze how natural selection has diversified its versatile RNA fold, we determined the X-ray crystal structure of the Burkholderia sp. TJI49 ykkC -I subtype-1 (Guanidine-I) riboswitch aptamer domain. Differing from the previously reported structures of orthologs from Dickeya dadantii and Sulfobacillus acidophilus , our Burkholderia structure reveals a chelated K + ion adjacent to two Mg 2+ ions in the guanidine-binding pocket. Thermal melting analysis shows that K + chelation, which induces localized conformational changes in the binding pocket, improves guanidinium-RNA interactions. Analysis of ribosome structures suggests that the [K + (Mg 2+ ) 2 ] ion triad is uncommon. It is, however, reminiscent of metal ion clusters found in the active sites of ribozymes and DNA polymerases. Previous structural characterization of ykkC -I subtype-2 RNAs, which bind the effector ligands ppGpp and PRPP, indicate that in those paralogs, an adenine responsible for K + chelation in the Burkholderia Guanidine-I riboswitch is replaced by a pyrimidine. This mutation results in a water molecule and Mg 2+ ion binding in place of the K + ion. Thus, our structural analysis demonstrates how ion and solvent chelation tune divergent ligand specificity and affinity among ykkC -I riboswitches.
- Biochemistry and Biophysics Center, National Heart, Lung, and Blood Institute, Bethesda, Maryland 20892-8012, USA.
Organizational Affiliation: