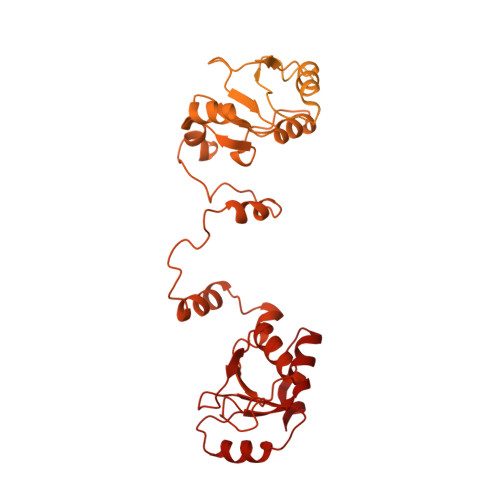
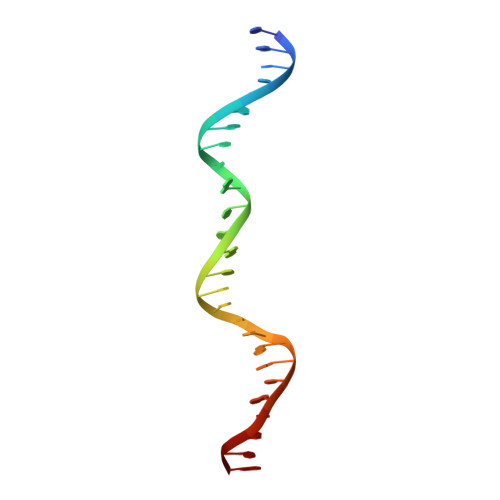
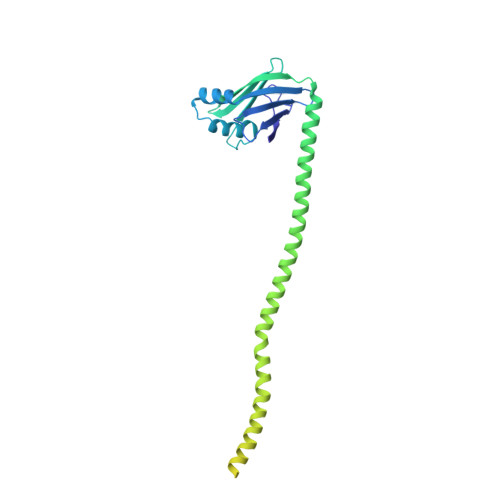
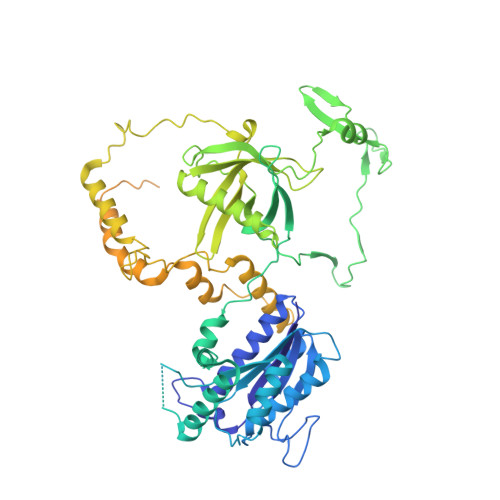
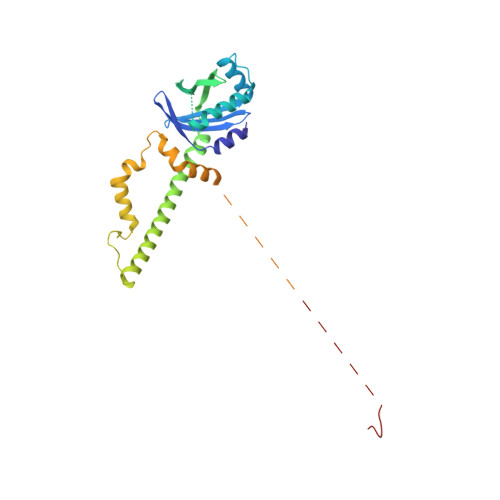
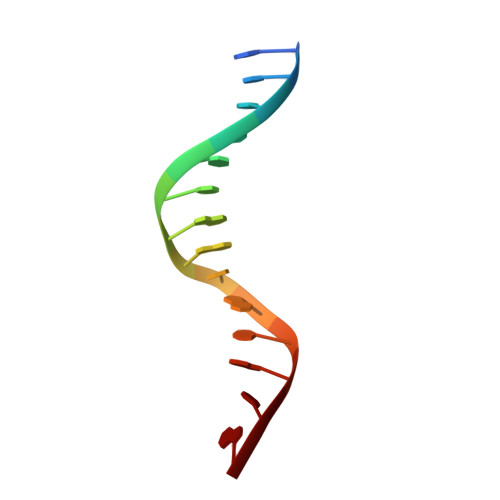
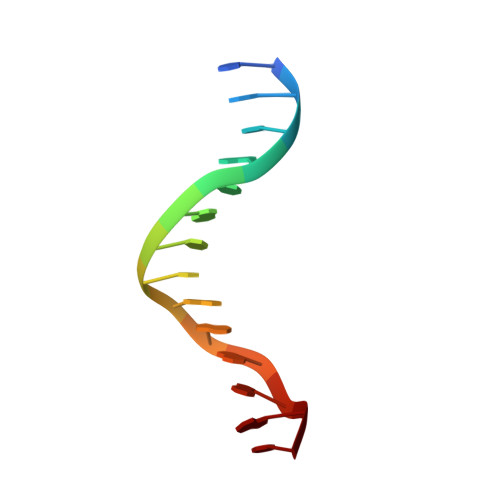
Structural basis of long-range to short-range synaptic transition in NHEJ.
Chen, S., Lee, L., Naila, T., Fishbain, S., Wang, A., Tomkinson, A.E., Lees-Miller, S.P., He, Y.(2021) Nature 593: 294-298
- PubMed: 33854234 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03458-7
- Primary Citation Related Structures:
7LSY, 7LT3 - PubMed Abstract:
DNA double-strand breaks (DSBs) are a highly cytotoxic form of DNA damage and the incorrect repair of DSBs is linked to carcinogenesis 1,2 . The conserved error-prone non-homologous end joining (NHEJ) pathway has a key role in determining the effects of DSB-inducing agents that are used to treat cancer as well as the generation of the diversity in antibodies and T cell receptors 2,3 . Here we applied single-particle cryo-electron microscopy to visualize two key DNA-protein complexes that are formed by human NHEJ factors. The Ku70/80 heterodimer (Ku), the catalytic subunit of the DNA-dependent protein kinase (DNA-PKcs), DNA ligase IV (LigIV), XRCC4 and XLF form a long-range synaptic complex, in which the DNA ends are held approximately 115 Å apart. Two DNA end-bound subcomplexes comprising Ku and DNA-PKcs are linked by interactions between the DNA-PKcs subunits and a scaffold comprising LigIV, XRCC4, XLF, XRCC4 and LigIV. The relative orientation of the DNA-PKcs molecules suggests a mechanism for autophosphorylation in trans, which leads to the dissociation of DNA-PKcs and the transition into the short-range synaptic complex. Within this complex, the Ku-bound DNA ends are aligned for processing and ligation by the XLF-anchored scaffold, and a single catalytic domain of LigIV is stably associated with a nick between the two Ku molecules, which suggests that the joining of both strands of a DSB involves both LigIV molecules.
- Department of Molecular Biosciences, Northwestern University, Evanston, IL, USA.
Organizational Affiliation: