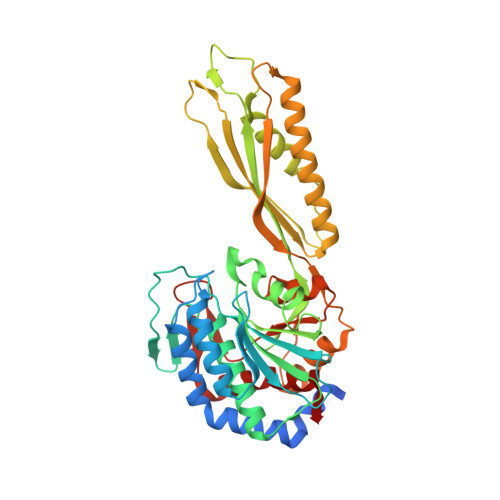
Sphingomonas sp. KT-1 PahZ2 Structure Reveals a Role for Conformational Dynamics in Peptide Bond Hydrolysis.
Brambley, C.A., Yared, T.J., Gonzalez, M., Jansch, A.L., Wallen, J.R., Weiland, M.H., Miller, J.M.(2021) J Phys Chem B 125: 5722-5739
- PubMed: 34060838
- DOI: https://doi.org/10.1021/acs.jpcb.1c01216
- Primary Citation Related Structures:
7LJH, 7LJI - PubMed Abstract:
Poly(aspartic acid) (PAA) is a common water-soluble polycarboxylate used in a broad range of applications. PAA biodegradation and environmental assimilation were first identified in river water bacterial strains, Sphingomonas sp. KT-1 and Pedobacter sp. KP-2. Within Sphingomonas sp. KT-1, PahZ1 KT-1 cleaves β-amide linkages to oligo(aspartic acid) and then is degraded by PahZ2 KT-1 . Recently, we reported the first structure for PahZ1 KT-1 . Here, we report novel structures for PahZ2 KT-1 bound to either Gd 3+ /Sm 3+ or Zn 2+ cations in a dimeric state consistent with M28 metallopeptidase family members. PahZ2 KT-1 monomers include a dimerization domain and a catalytic domain with dual Zn 2+ cations. MD methods predict the putative substrate binding site to span across the dimerization and catalytic domains, where NaCl promotes the transition from an open conformation to a closed conformation that positions the substrate adjacent to catalytic zinc ions. Structural knowledge of PahZ1 KT-1 and PahZ2 KT-1 will allow for protein engineering endeavors to develop novel biodegradation reagents.
- Department of Chemistry, Middle Tennessee State University, 1301 East Main Street, Murfreesboro, Tennessee 37132, United States.
Organizational Affiliation: