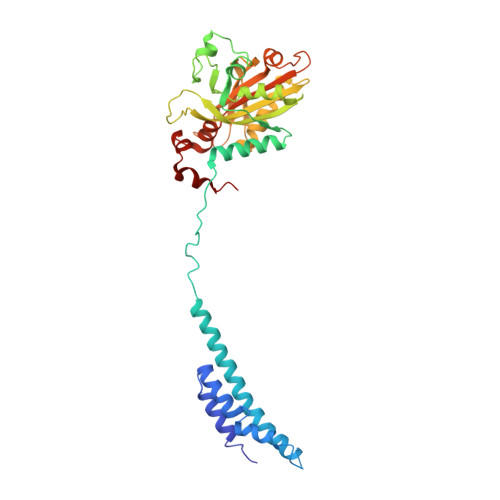
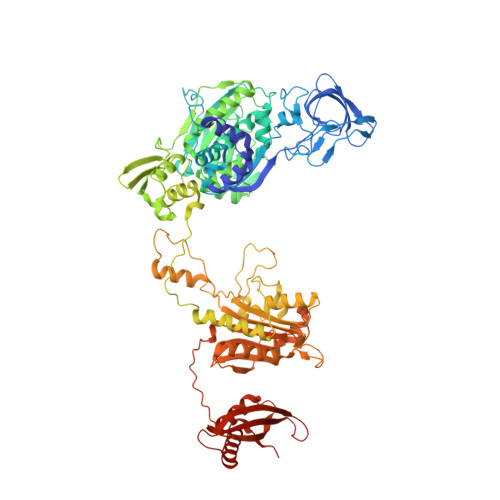
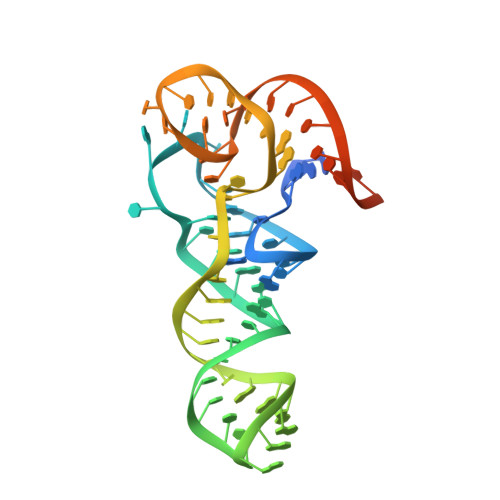
Mycobacterium tuberculosis Phe-tRNA synthetase: structural insights into tRNA recognition and aminoacylation.
Michalska, K., Jedrzejczak, R., Wower, J., Chang, C., Baragana, B., Gilbert, I.H., Forte, B., Joachimiak, A.(2021) Nucleic Acids Res 49: 5351-5368
- PubMed: 33885823 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab272
- Primary Citation Related Structures:
7K98, 7K9M, 7KA0, 7KAB - PubMed Abstract:
Tuberculosis, caused by Mycobacterium tuberculosis, responsible for ∼1.5 million fatalities in 2018, is the deadliest infectious disease. Global spread of multidrug resistant strains is a public health threat, requiring new treatments. Aminoacyl-tRNA synthetases are plausible candidates as potential drug targets, because they play an essential role in translating the DNA code into protein sequence by attaching a specific amino acid to their cognate tRNAs. We report structures of M. tuberculosis Phe-tRNA synthetase complexed with an unmodified tRNAPhe transcript and either L-Phe or a nonhydrolyzable phenylalanine adenylate analog. High-resolution models reveal details of two modes of tRNA interaction with the enzyme: an initial recognition via indirect readout of anticodon stem-loop and aminoacylation ready state involving interactions of the 3' end of tRNAPhe with the adenylate site. For the first time, we observe the protein gate controlling access to the active site and detailed geometry of the acyl donor and tRNA acceptor consistent with accepted mechanism. We biochemically validated the inhibitory potency of the adenylate analog and provide the most complete view of the Phe-tRNA synthetase/tRNAPhe system to date. The presented topography of amino adenylate-binding and editing sites at different stages of tRNA binding to the enzyme provide insights for the rational design of anti-tuberculosis drugs.
- Center for Structural Genomics of Infectious Diseases, Consortium for Advanced Science and Engineering, University of Chicago, Chicago, IL 60667, USA.
Organizational Affiliation: