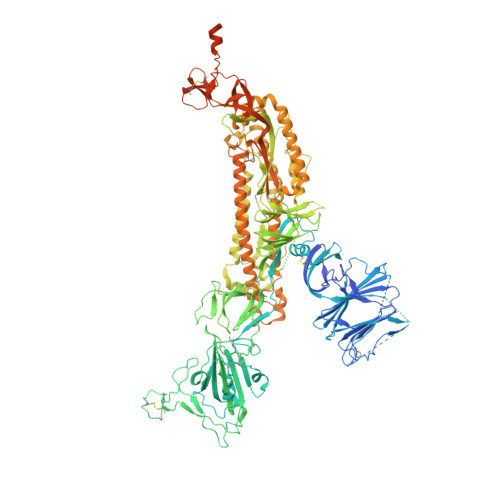
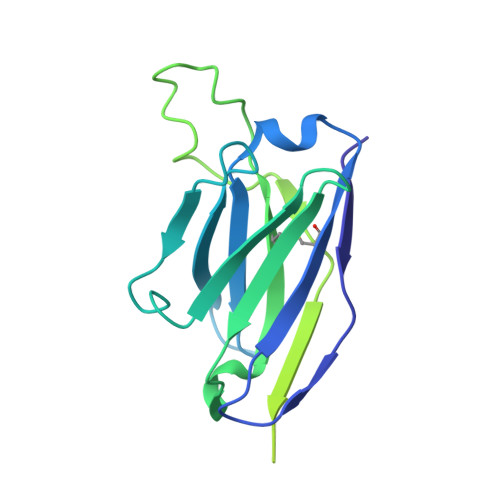
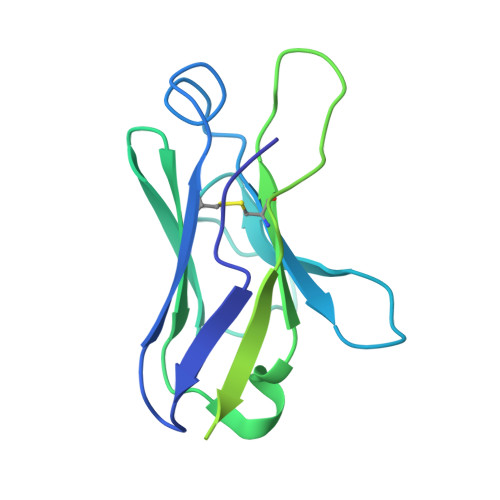
SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies.
Barnes, C.O., Jette, C.A., Abernathy, M.E., Dam, K.A., Esswein, S.R., Gristick, H.B., Malyutin, A.G., Sharaf, N.G., Huey-Tubman, K.E., Lee, Y.E., Robbiani, D.F., Nussenzweig, M.C., West Jr., A.P., Bjorkman, P.J.(2020) Nature 588: 682-687
- PubMed: 33045718 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-020-2852-1
- Primary Citation Related Structures:
7K8M, 7K8N, 7K8O, 7K8P, 7K8Q, 7K8R, 7K8S, 7K8T, 7K8U, 7K8V, 7K8W, 7K8X, 7K8Y, 7K8Z, 7K90 - PubMed Abstract:
The coronavirus disease 2019 (COVID-19) pandemic presents an urgent health crisis. Human neutralizing antibodies that target the host ACE2 receptor-binding domain (RBD) of the severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) spike protein 1-5 show promise therapeutically and are being evaluated clinically 6-8 . Here, to identify the structural correlates of SARS-CoV-2 neutralization, we solved eight new structures of distinct COVID-19 human neutralizing antibodies 5 in complex with the SARS-CoV-2 spike trimer or RBD. Structural comparisons allowed us to classify the antibodies into categories: (1) neutralizing antibodies encoded by the VH3-53 gene segment with short CDRH3 loops that block ACE2 and bind only to 'up' RBDs; (2) ACE2-blocking neutralizing antibodies that bind both up and 'down' RBDs and can contact adjacent RBDs; (3) neutralizing antibodies that bind outside the ACE2 site and recognize both up and down RBDs; and (4) previously described antibodies that do not block ACE2 and bind only to up RBDs 9 . Class 2 contained four neutralizing antibodies with epitopes that bridged RBDs, including a VH3-53 antibody that used a long CDRH3 with a hydrophobic tip to bridge between adjacent down RBDs, thereby locking the spike into a closed conformation. Epitope and paratope mapping revealed few interactions with host-derived N-glycans and minor contributions of antibody somatic hypermutations to epitope contacts. Affinity measurements and mapping of naturally occurring and in vitro-selected spike mutants in 3D provided insight into the potential for SARS-CoV-2 to escape from antibodies elicited during infection or delivered therapeutically. These classifications and structural analyses provide rules for assigning current and future human RBD-targeting antibodies into classes, evaluating avidity effects and suggesting combinations for clinical use, and provide insight into immune responses against SARS-CoV-2.
- Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, CA, USA.
Organizational Affiliation: