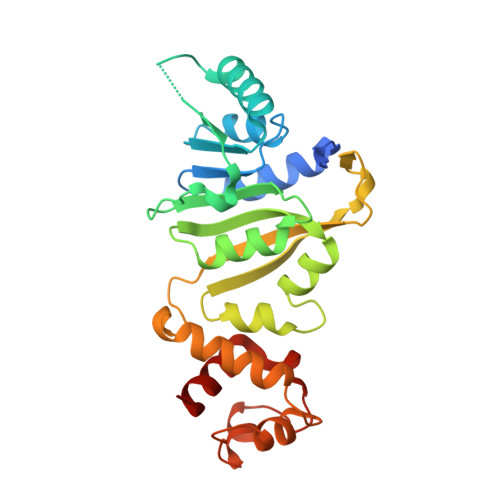
Crystal structure and functional analysis of mycobacterial erythromycin resistance methyltransferase Erm38 reveals its RNA-binding site.
Goh, B.C., Xiang, X., Lescar, J., Dedon, P.C.(2022) J Biological Chem 298: 101571-101571
- PubMed: 35007529
- DOI: https://doi.org/10.1016/j.jbc.2022.101571
- Primary Citation Related Structures:
7F8A, 7F8B, 7F8C - PubMed Abstract:
Erythromycin resistance methyltransferases (Erms) confer resistance to macrolide, lincosamide, and streptogramin antibiotics in Gram-positive bacteria and mycobacteria. Although structural information for ErmAM, ErmC, and ErmE exists from Gram-positive bacteria, little is known about the Erms in mycobacteria, as there are limited biochemical data and no structures available. Here, we present crystal structures of Erm38 from Mycobacterium smegmatis in apoprotein and cofactor-bound forms. Based on structural analysis and mutagenesis, we identified several catalytically critical, positively charged residues at a putative RNA-binding site. We found that mutation of any of these sites is sufficient to abolish methylation activity, whereas the corresponding RNA-binding affinity of Erm38 remains unchanged. The methylation reaction thus appears to require a precise ensemble of amino acids to accurately position the RNA substrate, such that the target nucleotide can be methylated. In addition, we computationally constructed a model of Erm38 in complex with a 32-mer RNA substrate. This model shows the RNA substrate stably bound to Erm38 by a patch of positively charged residues. Furthermore, a π-π stacking interaction between a key aromatic residue of Erm38 and a target adenine of the RNA substrate forms a critical interaction needed for methylation. Taken together, these data provide valuable insights into Erm-RNA interactions, which will aid subsequent structure-based drug design efforts.
- Antimicrobial Resistance Interdisciplinary Research Group, Singapore-MIT Alliance for Research and Technology Centre, Singapore, Singapore; NTU Institute of Structural Biology, Experimental Medicine Building (EMB), Nanyang Technological University, Singapore, Singapore. Electronic address: boonchong@smart.mit.edu.
Organizational Affiliation: