Isolation, characterization, and structure-based engineering of a neutralizing nanobody against SARS-CoV-2.
Li, T., Zhou, B., Li, Y., Huang, S., Luo, Z., Zhou, Y., Lai, Y., Gautam, A., Bourgeau, S., Wang, S., Bao, J., Tan, J., Lavillette, D., Li, D.(2022) Int J Biol Macromol 209: 1379-1388
- PubMed: 35460753 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.ijbiomac.2022.04.096
- Primary Citation Related Structures:
7F5G - PubMed Abstract:
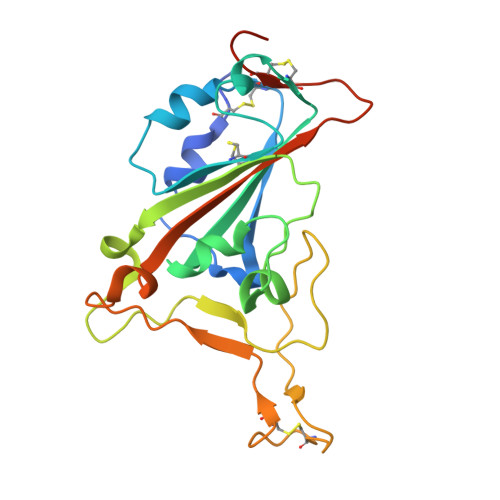
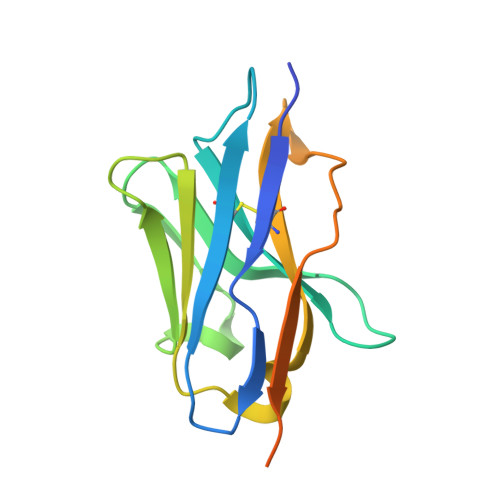
SARS-CoV-2 engages with human cells through the binding of its Spike receptor-binding domain (S-RBD) to the receptor ACE2. Molecular blocking of this engagement represents a proven strategy to treat COVID-19. Here, we report a single-chain antibody (nanobody, DL4) isolated from immunized alpaca with picomolar affinity to RBD. DL4 neutralizes SARS-CoV-2 pseudoviruses with an IC 50 of 0.101 μg mL -1 (6.2 nM). A crystal structure of the DL4-RBD complex at 1.75-Å resolution unveils the interaction detail and reveals a direct competition mechanism for DL4's ACE2-blocking and hence neutralizing activity. The structural information allows us to rationally design a mutant with higher potency. Our work adds diversity of neutralizing nanobodies against SARS-CoV-2 and should encourage protein engineering to improve antibody affinities in general.
- State Key Laboratory of Molecular Biology, CAS Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences (CAS), 320 Yueyang Road, Shanghai 200030, China.
Organizational Affiliation: