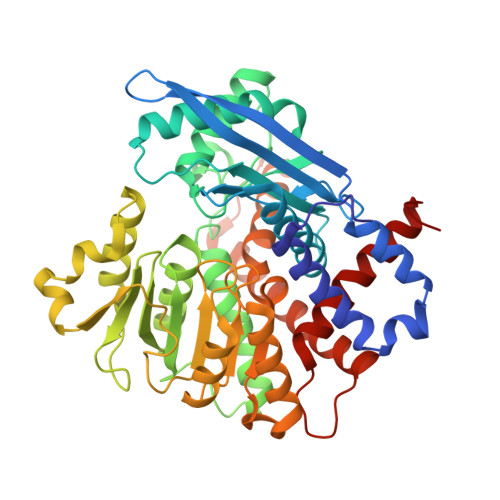
Molecular insights into the inhibition of glutamate dehydrogenase by the dicarboxylic acid metabolites.
Godsora, B.K.J., Prakash, P., Punekar, N.S., Bhaumik, P.(2022) Proteins 90: 810-823
- PubMed: 34748226 Search on PubMed
- DOI: https://doi.org/10.1002/prot.26276
- Primary Citation Related Structures:
7ECR, 7ECS, 7ECT - PubMed Abstract:
Glutamate dehydrogenase (GDH) is a salient metabolic enzyme which catalyzes the NAD + - or NADP + -dependent reversible conversion of α-ketoglutarate (AKG) to l-glutamate; and thereby connects the carbon and nitrogen metabolism cycles in all living organisms. The function of GDH is extensively regulated by both metabolites (citrate, succinate, etc.) and non-metabolites (ATP, NADH, etc.) but sufficient molecular evidences are lacking to rationalize the inhibitory effects by the metabolites. We have expressed and purified NADP + -dependent Aspergillus terreus GDH (AtGDH) in recombinant form. Succinate, malonate, maleate, fumarate, and tartrate independently inhibit the activity of AtGDH to different extents. The crystal structures of AtGDH complexed with the dicarboxylic acid metabolites and the coenzyme NADPH have been determined. Although AtGDH structures are not complexed with substrate; surprisingly, they acquire super closed conformation like previously reported for substrate and coenzyme bound catalytically competent Aspergillus niger GDH (AnGDH). These dicarboxylic acid metabolites partially occupy the same binding pocket as substrate; but interact with varying polar interactions and the coenzyme NADPH binds to the Domain-II of AtGDH. The low inhibition potential of tartrate as compared to other dicarboxylic acid metabolites is due to its weaker interactions of carboxylate groups with AtGDH. Our results suggest that the length of carbon skeleton and positioning of the carboxylate groups of inhibitors between two conserved lysine residues at the GDH active site might be the determinants of their inhibitory potency. Molecular details on the dicarboxylic acid metabolites bound AtGDH active site architecture presented here would be applicable to GDHs in general.
- Department of Biosciences and Bioengineering, Indian Institute of Technology Bombay, Mumbai, Maharashtra, India.
Organizational Affiliation: