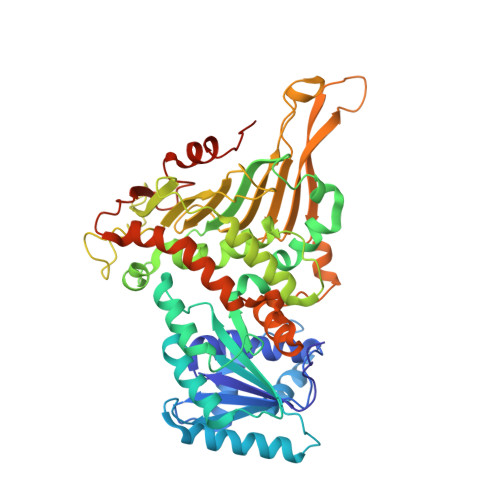
Structural basis for substrate recognition of glucose-6-phosphate dehydrogenase from Kluyveromyces lactis.
Vu, H.H., Jin, C., Chang, J.H.(2021) Biochem Biophys Res Commun 553: 85-91
- PubMed: 33765558
- DOI: https://doi.org/10.1016/j.bbrc.2021.02.088
- Primary Citation Related Structures:
7E6H, 7E6I - PubMed Abstract:
Glucose-6-phosphate dehydrogenase is the first enzyme in the pentose phosphate pathway. The reaction catalyzed by the enzyme is considered to be the main source of reducing power for nicotinamide adenine dinucleotide phosphate (NADPH) and is a precursor of 5-carbon sugar used by cells. To uncover the structural features of the enzyme, we determined the crystal structures of glucose-6-phosphate dehydrogenase from Kluyveromyces lactis (KlG6PD) in both the apo form and a binary complex with its substrate glucose-6-phosphate. KlG6PD contains a Rossman-like domain for cofactor NADPH binding; it also presents a typical antiparallel β sheet at the C-terminal domain with relatively the same pattern as those of other homologous structures. Moreover, our structural and biochemical analyses revealed that Lys153 contributes significantly to substrate G6P recognition. This study may provide insights into the structural variation and catalytic features of the G6PD enzyme.
- Department of Biology Education, Kyungpook National University, Daegu, 41566, South Korea.
Organizational Affiliation: