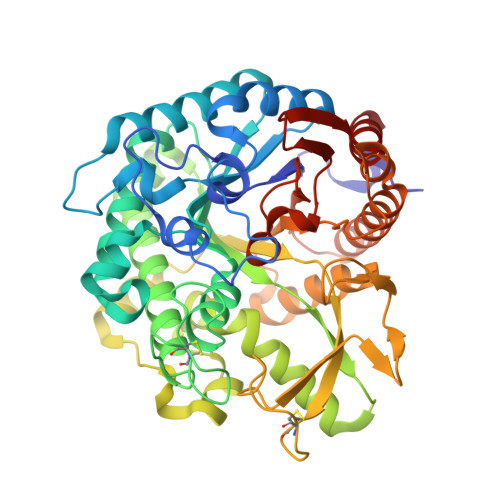
Structural analysis of rice Os4BGlu18 monolignol beta-glucosidase.
Baiya, S., Pengthaisong, S., Kitjaruwankul, S., Ketudat Cairns, J.R.(2021) PLoS One 16: e0241325-e0241325
- PubMed: 33471829 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0241325
- Primary Citation Related Structures:
7D6A, 7D6B - PubMed Abstract:
Monolignol glucosides are storage forms of monolignols, which are polymerized to lignin to strengthen plant cell walls. The conversion of monolignol glucosides to monolignols is catalyzed by monolignol β-glucosidases. Rice Os4BGlu18 β-glucosidase catalyzes hydrolysis of the monolignol glucosides, coniferin, syringin, and p-coumaryl alcohol glucoside more efficiently than other natural substrates. To understand more clearly the basis for substrate specificity of a monolignol β-glucosidase, the structure of Os4BGlu18 was determined by X-ray crystallography. Crystals of Os4BGlu18 and its complex with δ-gluconolactone diffracted to 1.7 and 2.1 Å resolution, respectively. Two protein molecules were found in the asymmetric unit of the P212121 space group of their isomorphous crystals. The Os4BGlu18 structure exhibited the typical (β/α)8 TIM barrel of glycoside hydrolase family 1 (GH1), but the four variable loops and two disulfide bonds appeared significantly different from other known structures of GH1 β-glucosidases. Molecular docking studies of the Os4BGlu18 structure with monolignol substrate ligands placed the glycone in a similar position to the δ-gluconolactone in the complex structure and revealed the interactions between protein and ligands. Molecular docking, multiple sequence alignment, and homology modeling identified amino acid residues at the aglycone-binding site involved in substrate specificity for monolignol β-glucosides. Thus, the structural basis of substrate recognition and hydrolysis by monolignol β-glucosidases was elucidated.
- Faculty of Science at Sriracha, Kasetsart University, Sriracha Campus, Sriracha, Chonburi, Thailand.
Organizational Affiliation: