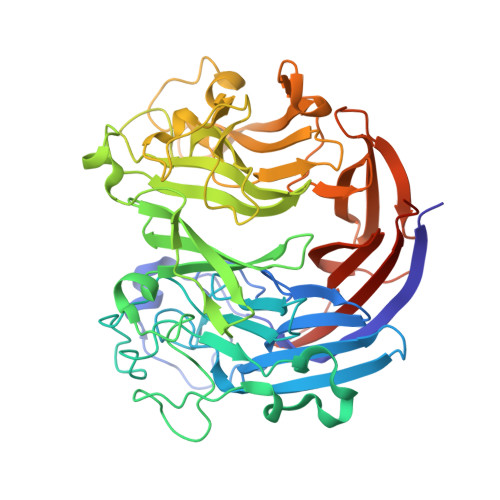
Mechanistic Insights into Substrate Recognition and Catalysis of a New Ulvan Lyase of Polysaccharide Lyase Family 24.
Xu, F., Dong, F., Sun, X.H., Cao, H.Y., Fu, H.H., Li, C.Y., Zhang, X.Y., McMinn, A., Zhang, Y.Z., Wang, P., Chen, X.L.(2021) Appl Environ Microbiol 87: e0041221-e0041221
- PubMed: 33771786
- DOI: https://doi.org/10.1128/AEM.00412-21
- Primary Citation Related Structures:
7CZH, 7DRQ - PubMed Abstract:
Ulvan is an important marine polysaccharide. Bacterial ulvan lyases play important roles in ulvan degradation and marine carbon cycling. Until now, only a small number of ulvan lyases have been characterized. Here, a new ulvan lyase, Uly1, belonging to polysaccharide lyase family 24 (PL24) from the marine bacterium Catenovulum maritimum, is characterized. The optimal temperature and pH for Uly1 to degrade ulvan are 40°C and pH 9.0, respectively. Uly1 degrades ulvan polysaccharides in the endolytic manner, mainly producing ΔRha3S, consisting of an unsaturated 4-deoxy-l- threo -hex-4-enopyranosiduronic acid and a 3-O-sulfated α-l-rhamnose. The structure of Uly1 was resolved at a 2.10-Å resolution. Uly1 adopts a seven-bladed β-propeller architecture. Structural and site-directed mutagenesis analyses indicate that four highly conserved residues, H128, H149, Y223, and R239, are essential for catalysis. H128 functions as both the catalytic acid and base, H149 and R239 function as the neutralizers, and Y223 plays a supporting role in catalysis. Structural comparison and sequence alignment suggest that Uly1 and many other PL24 enzymes may directly bind the substrate near the catalytic residues for catalysis, different from the PL24 ulvan lyase LOR_107, which adopts a two-stage substrate binding process. This study provides new insights into ulvan lyases and ulvan degradation. IMPORTANCE Ulvan is a major cell wall component of green algae of the genus Ulva. Many marine heterotrophic bacteria can produce extracellular ulvan lyases to degrade ulvan for a carbon nutrient. In addition, ulvan has a range of physiological bioactivities based on its specific chemical structure. Ulvan lyase thus plays an important role in marine carbon cycling and has great potential in biotechnological applications. However, only a small number of ulvan lyases have been characterized over the past 10 years. Here, based on biochemical and structural analyses, a new ulvan lyase of polysaccharide lyase family 24 is characterized, and its substrate recognition and catalytic mechanisms are revealed. Moreover, a new substrate binding process adopted by PL24 ulvan lyases is proposed. This study offers a better understanding of bacterial ulvan lyases and is helpful for studying the application potentials of ulvan lyases.
- State Key Laboratory of Microbial Technology, Shandong University, Qingdao, China.
Organizational Affiliation: