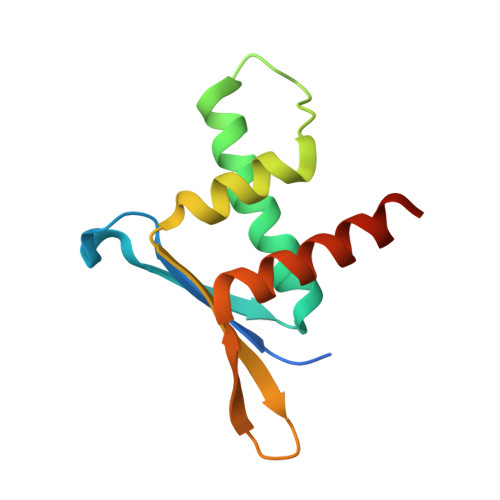
ERH facilitates microRNA maturation through the interaction with the N-terminus of DGCR8.
Kwon, S.C., Jang, H., Shen, S., Baek, S.C., Kim, K., Yang, J., Kim, J., Kim, J.S., Wang, S., Shi, Y., Li, F., Kim, V.N.(2020) Nucleic Acids Res 48: 11097-11112
- PubMed: 33035348
- DOI: https://doi.org/10.1093/nar/gkaa827
- Primary Citation Related Structures:
7CNC - PubMed Abstract:
The microprocessor complex cleaves the primary transcript of microRNA (pri-miRNA) to initiate miRNA maturation. Microprocessor is known to consist of RNase III DROSHA and dsRNA-binding DGCR8. Here, we identify Enhancer of Rudimentary Homolog (ERH) as a new component of Microprocessor. Through a crystal structure and biochemical experiments, we reveal that ERH uses its hydrophobic groove to bind to a conserved region in the N-terminus of DGCR8, in a 2:2 stoichiometry. Knock-down of ERH or deletion of the DGCR8 N-terminus results in a reduced processing of suboptimal pri-miRNAs in polycistronic miRNA clusters. ERH increases the processing of suboptimal pri-miR-451 in a manner dependent on its neighboring pri-miR-144. Thus, the ERH dimer may mediate 'cluster assistance' in which Microprocessor is loaded onto a poor substrate with help from a high-affinity substrate in the same cluster. Our study reveals a role of ERH in the miRNA biogenesis pathway.
- Center for RNA Research, Institute for Basic Science, Seoul 08826, Korea.
Organizational Affiliation: