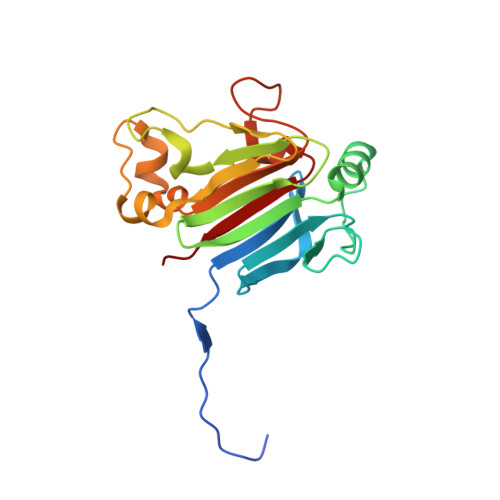
Structural basis for inhibitory effects of Smad7 on TGF-beta family signaling.
Murayama, K., Kato-Murayama, M., Itoh, Y., Miyazono, K., Miyazawa, K., Shirouzu, M.(2020) J Struct Biol 212: 107661-107661
- PubMed: 33166654 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2020.107661
- Primary Citation Related Structures:
7CD1 - PubMed Abstract:
Smad6 and Smad7 are classified as inhibitory Smads (I-Smads). They are crucial in the fine-tuning of signals by cytokines of the transforming growth factor-β (TGF-β) family. They are negative feedback regulators and principally target the activated type I receptors as well as the activated Smad complexes, but with distinct specificities. Smad7 inhibits Smad signaling from all seven type I receptors of the TGF-β family, whereas Smad6 preferentially inhibits Smad signaling from the bone morphogenetic protein (BMP) type I receptors, BMPR1A and BMPR1B. The target specificities are attributed to the C-terminal MH2 domain. Notably, Smad7 utilizes two alternative molecular surfaces for its inhibitory function against type I receptors. One is a basic groove composed of the first α-helix and the L3 loop, a structure that is shared with Smad6 and receptor-regulated Smads (R-Smads). The other is a three-finger-like structure (consisting of residues 331-361, 379-387, and the L3 loop) that is unique to Smad7. The underlying structural basis remains to be elucidated in detail. Here, we report the crystal structure of the MH2 domain of mouse Smad7 at 1.9 Å resolution. The three-finger-like structure is stabilized by a network of hydrogen bonds between residues 331-361 and 379-387, thus forming a molecular surface unique to Smad7. Furthermore, we discuss how Smad7 antagonizes the activated Smad complexes composed of R-Smad and Smad4, a common partner Smad.
- Laboratory for Protein Functional and Structural Biology, RIKEN Center for Biosystems Dynamics Research, Yokohama, 1-7-22 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan; Graduate School of Biomedical Engineering, Tohoku University, 2-1 Seiryomachi, Aoba, Sendai 980-8575, Japan.
Organizational Affiliation: