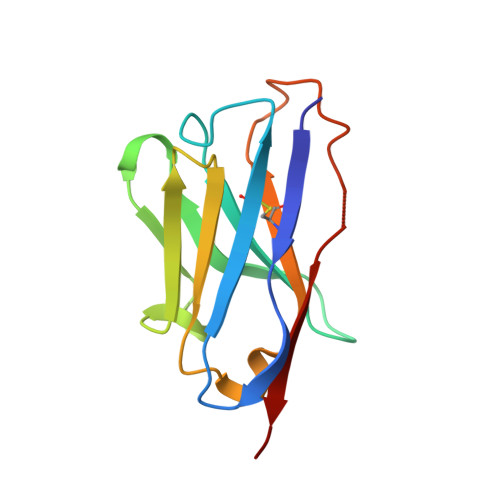
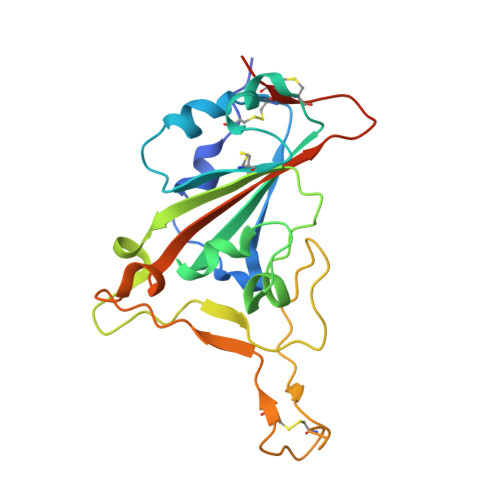
A synthetic nanobody targeting RBD protects hamsters from SARS-CoV-2 infection.
Li, T., Cai, H., Yao, H., Zhou, B., Zhang, N., van Vlissingen, M.F., Kuiken, T., Han, W., GeurtsvanKessel, C.H., Gong, Y., Zhao, Y., Shen, Q., Qin, W., Tian, X.X., Peng, C., Lai, Y., Wang, Y., Hutter, C.A.J., Kuo, S.M., Bao, J., Liu, C., Wang, Y., Richard, A.S., Raoul, H., Lan, J., Seeger, M.A., Cong, Y., Rockx, B., Wong, G., Bi, Y., Lavillette, D., Li, D.(2021) Nat Commun 12: 4635-4635
- PubMed: 34330908
- DOI: https://doi.org/10.1038/s41467-021-24905-z
- Primary Citation Related Structures:
7C8V, 7C8W, 7CAN - PubMed Abstract:
SARS-CoV-2, the causative agent of COVID-19 1 , features a receptor-binding domain (RBD) for binding to the host cell ACE2 protein 1-6 . Neutralizing antibodies that block RBD-ACE2 interaction are candidates for the development of targeted therapeutics 7-17 . Llama-derived single-domain antibodies (nanobodies, ~15 kDa) offer advantages in bioavailability, amenability, and production and storage owing to their small sizes and high stability. Here, we report the rapid selection of 99 synthetic nanobodies (sybodies) against RBD by in vitro selection using three libraries. The best sybody, MR3 binds to RBD with high affinity (K D = 1.0 nM) and displays high neutralization activity against SARS-CoV-2 pseudoviruses (IC 50 = 0.42 μg mL -1 ). Structural, biochemical, and biological characterization suggests a common neutralizing mechanism, in which the RBD-ACE2 interaction is competitively inhibited by sybodies. Various forms of sybodies with improved potency have been generated by structure-based design, biparatopic construction, and divalent engineering. Two divalent forms of MR3 protect hamsters from clinical signs after live virus challenge and a single dose of the Fc-fusion construct of MR3 reduces viral RNA load by 6 Log 10 . Our results pave the way for the development of therapeutic nanobodies against COVID-19 and present a strategy for rapid development of targeted medical interventions during an outbreak.
- State Key Laboratory of Molecular Biology, CAS Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences (CAS), Shanghai, China.
Organizational Affiliation: