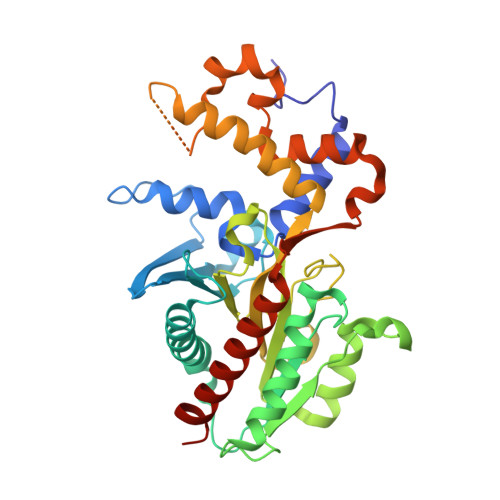
Structural insights into the psychrophilic germinal protease PaGPR and its autoinhibitory loop.
Lee, C.W., Lee, S., Jeong, C.S., Hwang, J., Chang, J.H., Choi, I.G., Kim, T.D., Park, H., Kim, H.Y., Lee, J.H.(2020) J Microbiol 58: 772-779
- PubMed: 32870483 Search on PubMed
- DOI: https://doi.org/10.1007/s12275-020-0292-0
- Primary Citation Related Structures:
7C4X - PubMed Abstract:
In spore forming microbes, germination protease (GPR) plays a key role in the initiation of the germination process. A critical step during germination is the degradation of small acid-soluble proteins (SASPs), which protect spore DNA from external stresses (UV, heat, low temperature, etc.). Inactive zymogen GPR can be activated by autoprocessing of the N-terminal pro-sequence domain. Activated GPR initiates the degradation of SASPs; however, the detailed mechanisms underlying the activation, catalysis, regulation, and substrate recognition of GPR remain elusive. In this study, we determined the crystal structure of GPR from Paenisporosarcina sp. TG-20 (PaGPR) in its inactive form at a resolution of 2.5 A. Structural analysis showed that the active site of PaGPR is sterically occluded by an inhibitory loop region (residues 202-216). The N-terminal region interacts directly with the self-inhibitory loop region, suggesting that the removal of the N-terminal pro-sequence induces conformational changes, which lead to the release of the self-inhibitory loop region from the active site. In addition, comparative sequence and structural analyses revealed that PaGPR contains two highly conserved Asp residues (D123 and D182) in the active site, similar to the putative aspartic acid protease GPR from Bacillus megaterium. The catalytic domain structure of PaGPR also shares similarities with the sequentially non-homologous proteins HycI and HybD. HycI and HybD are metal-loproteases that also contain two Asp (or Glu) residues in their active site, playing a role in metal binding. In summary, our results provide useful insights into the activation process of PaGPR and its active conformation.
- Unit of Research for Practical Application, Korea Polar Research Institute, Incheon, 21990, Republic of Korea.
Organizational Affiliation: