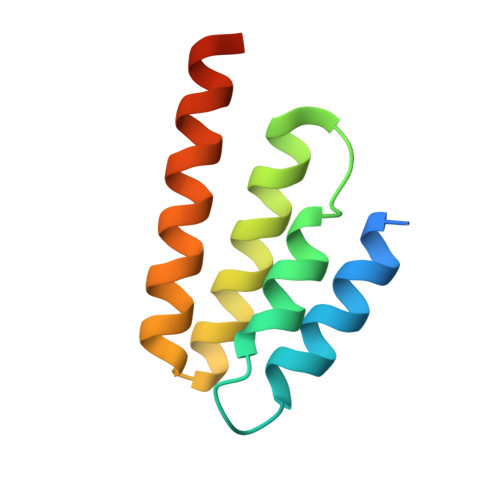
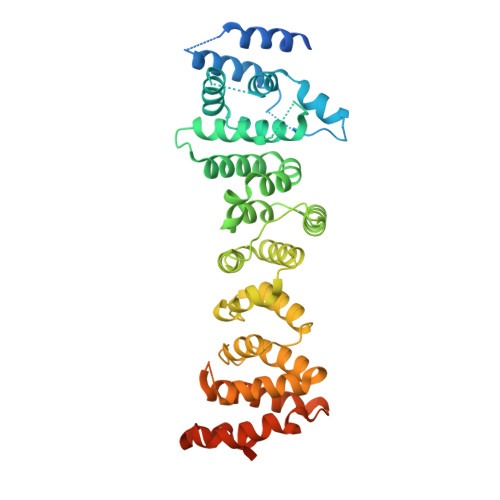Crystal structure of the INTS3/INTS6 complex reveals the functional importance of INTS3 dimerization in DSB repair.
Jia, Y., Cheng, Z., Bharath, S.R., Sun, Q., Su, N., Huang, J., Song, H.(2021) Cell Discov 7: 66-66
- PubMed: 34400606 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-021-00283-0
- Primary Citation Related Structures:
7BV7 - PubMed Abstract:
SOSS1 is a single-stranded DNA (ssDNA)-binding protein complex that plays a critical role in double-strand DNA break (DSB) repair. SOSS1 consists of three subunits: INTS3, SOSSC, and hSSB1, with INTS3 serving as a scaffold to stabilize this complex. Moreover, the integrator complex subunit 6 (INTS6) participates in the DNA damage response through direct binding to INTS3, but how INTS3 interacts with INTS6, thereby impacting DSB repair, is not clear. Here, we determined the crystal structure of the C-terminus of INTS3 (INTS3c) in complex with the C-terminus of INTS6 (INTS6c) at a resolution of 2.4 Å. Structural analysis revealed that two INTS3c subunits dimerize and interact with INTS6c via conserved residues. Subsequent biochemical analyses confirmed that INTS3c forms a stable dimer and INTS3 dimerization is important for recognizing the longer ssDNA. Perturbation of INTS3c dimerization and disruption of the INTS3c/INTS6c interaction impair the DSB repair process. Altogether, these results unravel the underappreciated role of INTS3 dimerization and the molecular basis of INTS3/INTS6 interaction in DSB repair.
- MOE Laboratory of Biosystems Homeostasis and Protection and Innovation Center for Cell Signaling Network, Life Sciences Institute, Zhejiang University, Hangzhou, Zhejiang, China.
Organizational Affiliation: