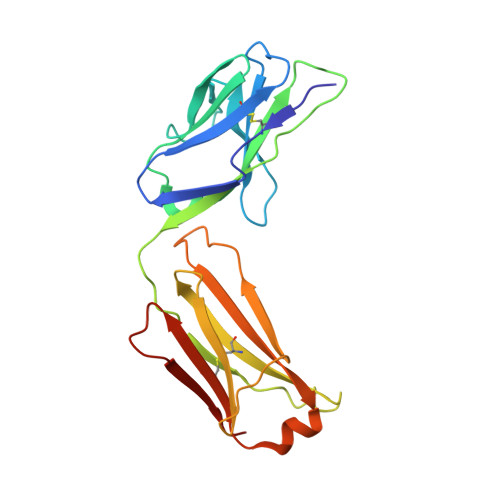
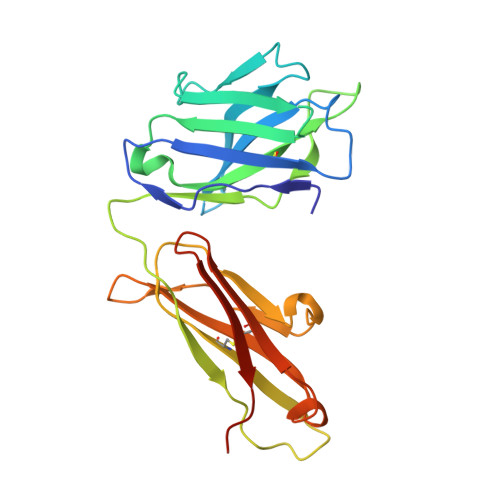
The role of membrane destabilisation and protein dynamics in BAM catalysed OMP folding.
White, P., Haysom, S.F., Iadanza, M.G., Higgins, A.J., Machin, J.M., Whitehouse, J.M., Horne, J.E., Schiffrin, B., Carpenter-Platt, C., Calabrese, A.N., Storek, K.M., Rutherford, S.T., Brockwell, D.J., Ranson, N.A., Radford, S.E.(2021) Nat Commun 12: 4174-4174
- PubMed: 34234105
- DOI: https://doi.org/10.1038/s41467-021-24432-x
- Primary Citation Related Structures:
7BM5, 7BNQ, 7NBX, 7NCS, 7ND0 - PubMed Abstract:
The folding of β-barrel outer membrane proteins (OMPs) in Gram-negative bacteria is catalysed by the β-barrel assembly machinery (BAM). How lateral opening in the β-barrel of the major subunit BamA assists in OMP folding, and the contribution of membrane disruption to BAM catalysis remain unresolved. Here, we use an anti-BamA monoclonal antibody fragment (Fab1) and two disulphide-crosslinked BAM variants (lid-locked (LL), and POTRA-5-locked (P5L)) to dissect these roles. Despite being lethal in vivo, we show that all complexes catalyse folding in vitro, albeit less efficiently than wild-type BAM. CryoEM reveals that while Fab1 and BAM-P5L trap an open-barrel state, BAM-LL contains a mixture of closed and contorted, partially-open structures. Finally, all three complexes globally destabilise the lipid bilayer, while BamA does not, revealing that the BAM lipoproteins are required for this function. Together the results provide insights into the role of BAM structure and lipid dynamics in OMP folding.
- Astbury Centre for Structural Molecular Biology, School of Molecular and Cellular Biology, Faculty of Biological Sciences, University of Leeds, Leeds, UK.
Organizational Affiliation: