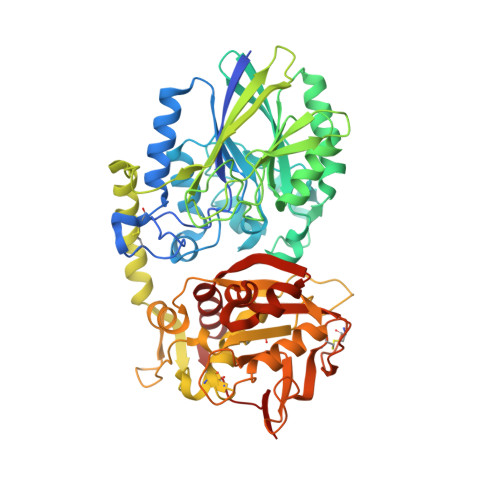
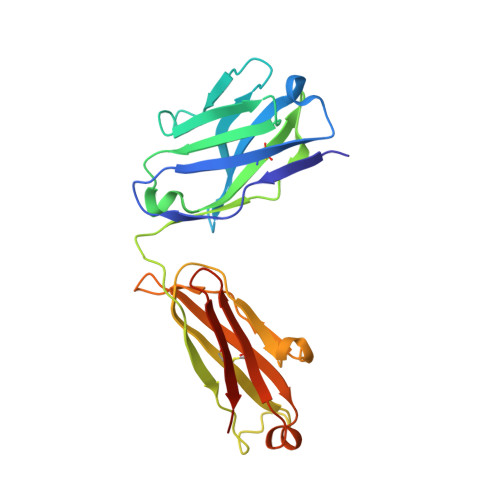
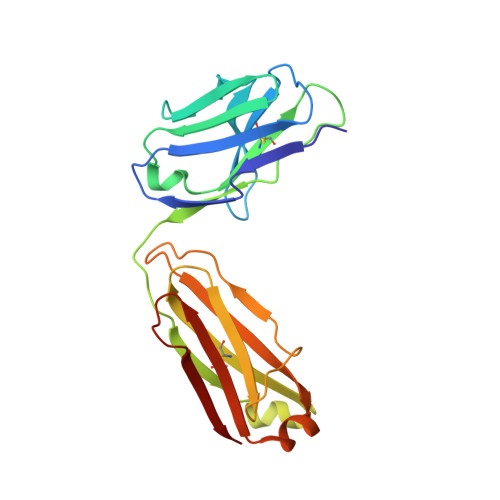
A Novel Antagonistic CD73 Antibody for Inhibition of the Immunosuppressive Adenosine Pathway.
Wurm, M., Schaaf, O., Reutner, K., Ganesan, R., Mostbock, S., Pelster, C., Bottcher, J., de Andrade Pereira, B., Taubert, C., Alt, I., Serna, G., Auguste, A., Stadermann, K.B., Delic, D., Han, F., Capdevila, J., Nuciforo, P.G., Kroe-Barrett, R., Adam, P.J., Vogt, A.B., Hofmann, I.(2021) Mol Cancer Ther 20: 2250-2261
- PubMed: 34482286 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1158/1535-7163.MCT-21-0107
- Primary Citation Related Structures:
7BBJ - PubMed Abstract:
Despite some impressive clinical results with immune checkpoint inhibitors, the majority of patients with cancer do not respond to these agents, in part due to immunosuppressive mechanisms in the tumor microenvironment. High levels of adenosine in tumors can suppress immune cell function, and strategies to target the pathway involved in its production have emerged. CD73 is a key enzyme involved in adenosine production. This led us to identify a novel humanized antagonistic CD73 antibody, mAb19, with distinct binding properties. mAb19 potently inhibits the enzymatic activity of CD73 in vitro , resulting in an inhibition of adenosine formation and enhanced T-cell activation. We then investigated the therapeutic potential of combining CD73 antagonism with other immune modulatory and chemotherapeutic agents. Combination of mAb19 with a PD-1 inhibitor increased T-cell activation in vitro Interestingly, this effect could be further enhanced with an agonist of the adenosine receptor ADORA3. Adenosine levels were found to be elevated upon doxorubicin treatment in vivo , which could be blocked by CD73 inhibition. Combining CD73 antagonism with doxorubicin resulted in superior responses in vivo Furthermore, a retrospective analysis of rectal cancer patient samples demonstrated an upregulation of the adenosine pathway upon chemoradiation, providing further rationale for combining CD73 inhibition with chemotherapeutic agents.This study demonstrates the ability of a novel CD73 antibody to enhance T-cell function through the potent suppression of adenosine levels. In addition, the data highlight combination opportunities with standard of care therapies as well as with an ADORA3 receptor agonist to treat patients with solid tumors.
- Boehringer Ingelheim RCV, GmbH & Co KG, Cancer Pharmacology and Disease Positioning, Vienna, Austria.
Organizational Affiliation: