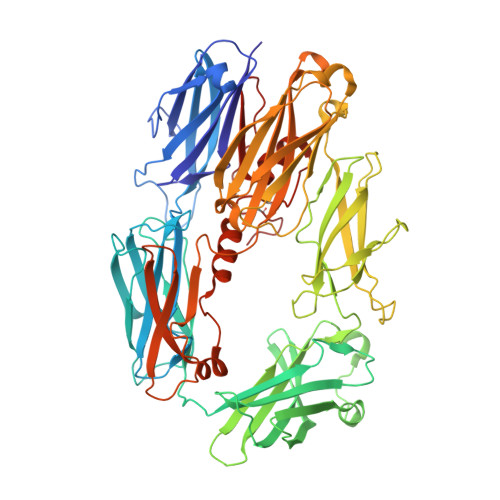
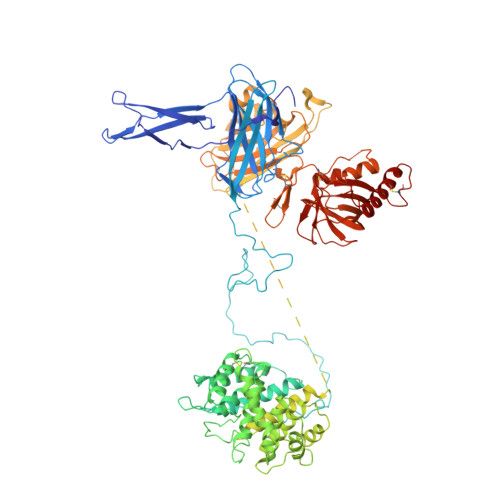
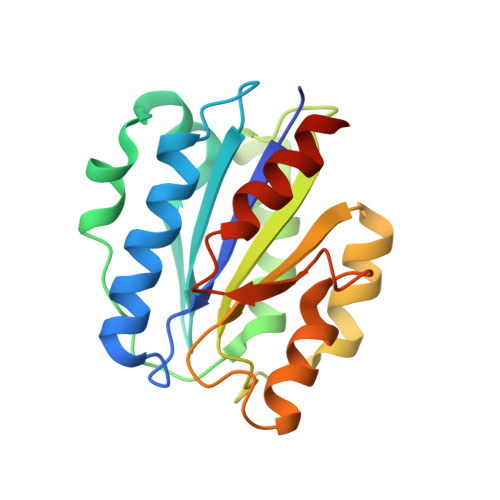
The crystal structure of iC3b-CR3 alpha I reveals a modular recognition of the main opsonin iC3b by the CR3 integrin receptor
Fernandez, F.J., Santos-Lopez, J., Martinez-Barricarte, R., Querol-Garcia, J., Martin-Merinero, H., Navas-Yuste, S., Savko, M., Shepard, W.E., Rodriguez de Cordoba, S., Vega, M.C.(2022) Nat Commun 13: 1955