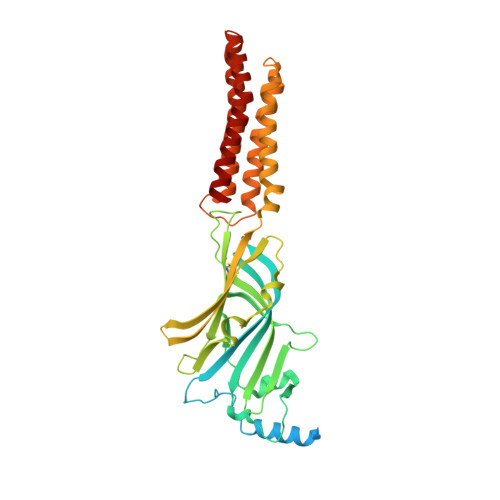
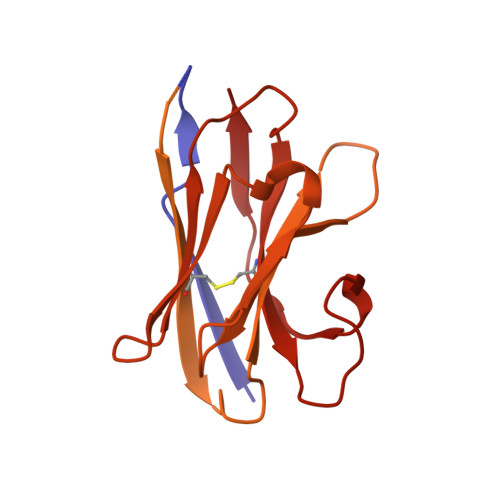
Single-particle cryo-EM at atomic resolution.
Nakane, T., Kotecha, A., Sente, A., McMullan, G., Masiulis, S., Brown, P.M.G.E., Grigoras, I.T., Malinauskaite, L., Malinauskas, T., Miehling, J., Uchanski, T., Yu, L., Karia, D., Pechnikova, E.V., de Jong, E., Keizer, J., Bischoff, M., McCormack, J., Tiemeijer, P., Hardwick, S.W., Chirgadze, D.Y., Murshudov, G., Aricescu, A.R., Scheres, S.H.W.(2020) Nature 587: 152-156
- PubMed: 33087931 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-020-2829-0
- Primary Citation Related Structures:
7A4M, 7A5V - PubMed Abstract:
The three-dimensional positions of atoms in protein molecules define their structure and their roles in biological processes. The more precisely atomic coordinates are determined, the more chemical information can be derived and the more mechanistic insights into protein function may be inferred. Electron cryo-microscopy (cryo-EM) single-particle analysis has yielded protein structures with increasing levels of detail in recent years 1,2 . However, it has proved difficult to obtain cryo-EM reconstructions with sufficient resolution to visualize individual atoms in proteins. Here we use a new electron source, energy filter and camera to obtain a 1.7 Å resolution cryo-EM reconstruction for a human membrane protein, the β3 GABA A receptor homopentamer 3 . Such maps allow a detailed understanding of small-molecule coordination, visualization of solvent molecules and alternative conformations for multiple amino acids, and unambiguous building of ordered acidic side chains and glycans. Applied to mouse apoferritin, our strategy led to a 1.22 Å resolution reconstruction that offers a genuine atomic-resolution view of a protein molecule using single-particle cryo-EM. Moreover, the scattering potential from many hydrogen atoms can be visualized in difference maps, allowing a direct analysis of hydrogen-bonding networks. Our technological advances, combined with further approaches to accelerate data acquisition and improve sample quality, provide a route towards routine application of cryo-EM in high-throughput screening of small molecule modulators and structure-based drug discovery.
- MRC Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: