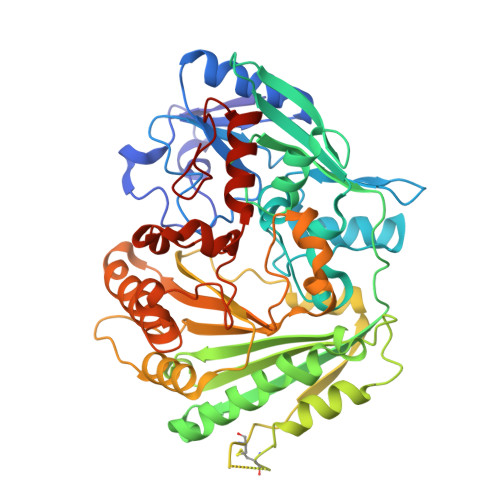
Structural Elucidation and Engineering of a Bacterial Carbohydrate Oxidase.
Boverio, A., Widodo, W.S., Santema, L.L., Rozeboom, H.J., Xiang, R., Guallar, V., Mattevi, A., Fraaije, M.W.(2023) Biochemistry 62: 429-436
- PubMed: 35881507 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.2c00307
- Primary Citation Related Structures:
7ZZK - PubMed Abstract:
Flavin-dependent carbohydrate oxidases are valuable tools in biotechnological applications due to their high selectivity in the oxidation of carbohydrates. In this study, we report the biochemical and structural characterization of a recently discovered carbohydrate oxidase from the bacterium Ralstonia solanacearum , which is a member of the vanillyl alcohol oxidase flavoprotein family. Due to its exceptionally high activity toward N -acetyl-d-galactosamine and N -acetyl-d-glucosamine, the enzyme was named N -acetyl-glucosamine oxidase (NagOx). In contrast to most known (fungal) carbohydrate oxidases, NagOx could be overexpressed in a bacterial host, which facilitated detailed biochemical and enzyme engineering studies. Steady state kinetic analyses revealed that non-acetylated hexoses were also accepted as substrates albeit with lower efficiency. Upon determination of the crystal structure, structural insights into NagOx were obtained. A large cavity containing a bicovalently bound FAD, tethered via histidyl and cysteinyl linkages, was observed. Substrate docking highlighted how a single residue (Leu251) plays a key role in the accommodation of N-acetylated sugars in the active site. Upon replacement of Leu251 (L251R mutant), an enzyme variant was generated with a drastically modified substrate acceptance profile, tuned toward non-N-acetylated monosaccharides and disaccharides. Furthermore, the activity toward bulkier substrates such as the trisaccharide maltotriose was introduced by this mutation. Due to its advantage of being overexpressed in a bacterial host, NagOx can be considered a promising alternative engineerable biocatalyst for selective oxidation of monosaccharides and oligosaccharides.
- Molecular Enzymology, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, 9747AG Groningen, The Netherlands.
Organizational Affiliation: