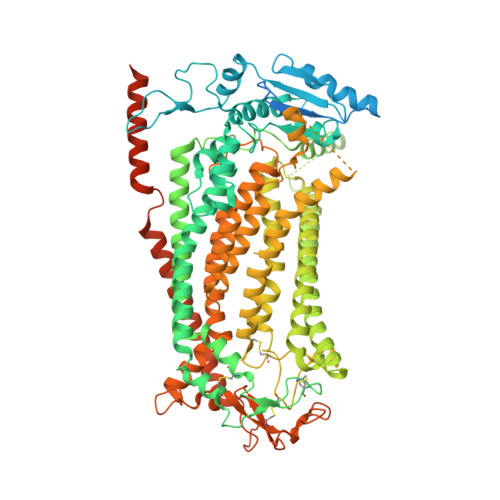
Inhibition mechanism of the chloride channel TMEM16A by the pore blocker 1PBC.
Lam, A.K.M., Rutz, S., Dutzler, R.(2022) Nat Commun 13: 2798-2798
- PubMed: 35589730 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30479-1
- Primary Citation Related Structures:
7ZK3 - PubMed Abstract:
TMEM16A, a calcium-activated chloride channel involved in multiple cellular processes, is a proposed target for diseases such as hypertension, asthma, and cystic fibrosis. Despite these therapeutic promises, its pharmacology remains poorly understood. Here, we present a cryo-EM structure of TMEM16A in complex with the channel blocker 1PBC and a detailed functional analysis of its inhibition mechanism. A pocket located external to the neck region of the hourglass-shaped pore is responsible for open-channel block by 1PBC and presumably also by its structural analogs. The binding of the blocker stabilizes an open-like conformation of the channel that involves a rearrangement of several pore helices. The expansion of the outer pore enhances blocker sensitivity and enables 1PBC to bind at a site within the transmembrane electric field. Our results define the mechanism of inhibition and gating and will facilitate the design of new, potent TMEM16A modulators.
- Department of Biochemistry, University of Zurich, Winterthurer Str. 190, CH-8057, Zurich, Switzerland. a.lam@bioc.uzh.ch.
Organizational Affiliation: