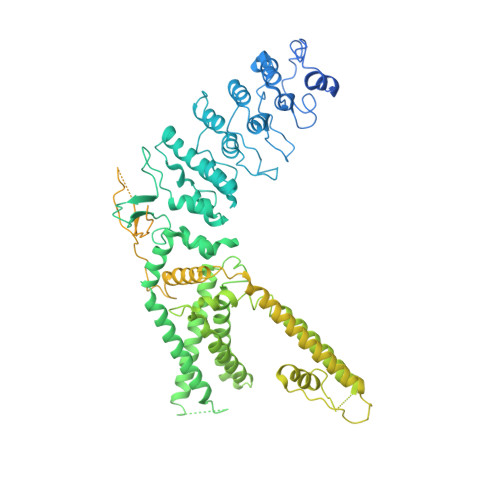
Cannabinoid non-cannabidiol site modulation of TRPV2 structure and function.
Zhang, L., Simonsen, C., Zimova, L., Wang, K., Moparthi, L., Gaudet, R., Ekoff, M., Nilsson, G., Hellmich, U.A., Vlachova, V., Gourdon, P., Zygmunt, P.M.(2022) Nat Commun 13: 7483-7483
- PubMed: 36470868 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-35163-y
- Primary Citation Related Structures:
7ZJD, 7ZJE, 7ZJG, 7ZJH, 7ZJI - PubMed Abstract:
TRPV2 is a ligand-operated temperature sensor with poorly defined pharmacology. Here, we combine calcium imaging and patch-clamp electrophysiology with cryo-electron microscopy (cryo-EM) to explore how TRPV2 activity is modulated by the phytocannabinoid Δ 9 -tetrahydrocannabiorcol (C16) and by probenecid. C16 and probenecid act in concert to stimulate TRPV2 responses including histamine release from rat and human mast cells. Each ligand causes distinct conformational changes in TRPV2 as revealed by cryo-EM. Although the binding for probenecid remains elusive, C16 associates within the vanilloid pocket. As such, the C16 binding location is distinct from that of cannabidiol, partially overlapping with the binding site of the TRPV2 inhibitor piperlongumine. Taken together, we discover a new cannabinoid binding site in TRPV2 that is under the influence of allosteric control by probenecid. This molecular insight into ligand modulation enhances our understanding of TRPV2 in normal and pathophysiology.
- Department of Clinical Sciences Malmö, Lund University, Malmö, Sweden.
Organizational Affiliation: