Gcf1p tetramerizes upon DNA binding, revealing a novel compaction method for mitochondrial DNA in Candida albicans.
Tarres-Sole, A., Ruiz-Lopez, E., Lyonnais, S., Sola, M.(2023) Nucleic Acids Res
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
(2023) Nucleic Acids Res
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Gcf1p | 245 | Candida albicans | Mutation(s): 0 Gene Names: GCF1, orf19.400, CAALFM_C108550CA |  | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q59QB8 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| 20-mer DNA | C [auth W], E [auth Y] | 20 | synthetic construct |  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 3 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| 20-mer DNA | D [auth X], F [auth Z] | 20 | synthetic construct |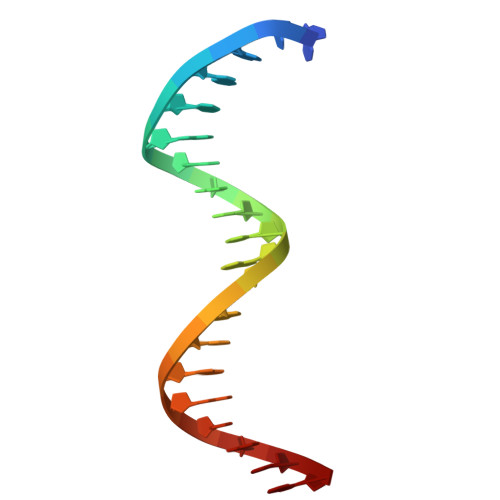  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GOL Download:Ideal Coordinates CCD File | G [auth A], H [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 66.63 | α = 90 |
| b = 96.9 | β = 90 |
| c = 113.06 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| MxCuBE | data collection |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Ministerio de Ciencia e Innovacion (MCIN) | Spain | RTI2018-101015-B-100 |