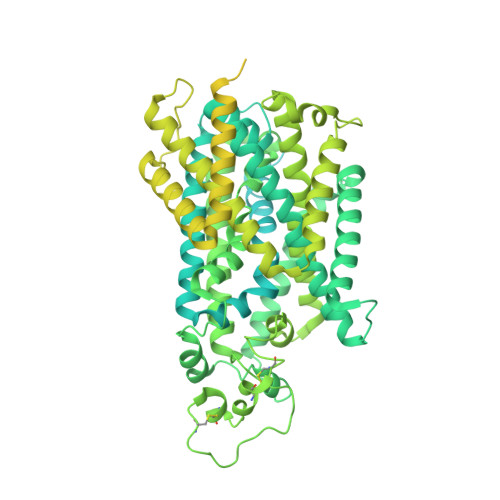
Cryo-EM structure of the human NKCC1 transporter reveals mechanisms of ion coupling and specificity.
Neumann, C., Rosenbaek, L.L., Flygaard, R.K., Habeck, M., Karlsen, J.L., Wang, Y., Lindorff-Larsen, K., Gad, H.H., Hartmann, R., Lyons, J.A., Fenton, R.A., Nissen, P.(2022) EMBO J 41: e110169-e110169
- PubMed: 36239040 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2021110169
- Primary Citation Related Structures:
7ZGO - PubMed Abstract:
The sodium-potassium-chloride transporter NKCC1 of the SLC12 family performs Na + -dependent Cl - - and K + -ion uptake across plasma membranes. NKCC1 is important for regulating cell volume, hearing, blood pressure, and regulation of hyperpolarizing GABAergic and glycinergic signaling in the central nervous system. Here, we present a 2.6 Å resolution cryo-electron microscopy structure of human NKCC1 in the substrate-loaded (Na + , K + , and 2 Cl - ) and occluded, inward-facing state that has also been observed for the SLC6-type transporters MhsT and LeuT. Cl - binding at the Cl1 site together with the nearby K + ion provides a crucial bridge between the LeuT-fold scaffold and bundle domains. Cl - -ion binding at the Cl2 site seems to undertake a structural role similar to conserved glutamate of SLC6 transporters and may allow for Cl - -sensitive regulation of transport. Supported by functional studies in mammalian cells and computational simulations, we describe a putative Na + release pathway along transmembrane helix 5 coupled to the Cl2 site. The results provide insight into the structure-function relationship of NKCC1 with broader implications for other SLC12 family members.
- Danish Research Institute of Translational Neuroscience-DANDRITE, Nordic EMBL Partnership for Molecular Medicine, Aarhus, Denmark.
Organizational Affiliation: