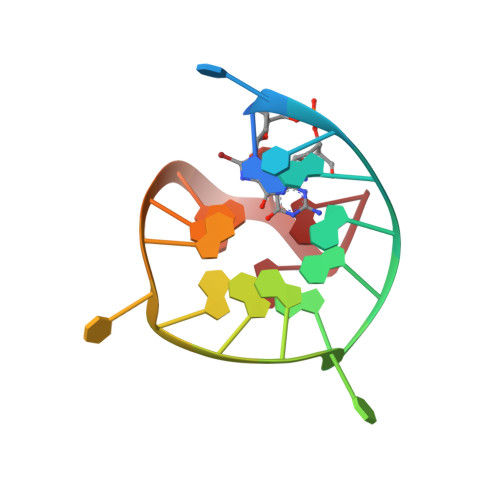
Guiding the folding of G-quadruplexes through loop residue interactions.
Jana, J., Vianney, Y.M., Schroder, N., Weisz, K.(2022) Nucleic Acids Res 50: 7161-7175
- PubMed: 35758626 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac549
- Primary Citation Related Structures:
7ZEK, 7ZEM, 7ZEO - PubMed Abstract:
A G-rich sequence was designed to allow folding into either a stable parallel or hybrid-type topology. With the parent sequence featuring coexisting species, various related sequences with single and double mutations and with a shortened central propeller loop affected the topological equilibrium. Two simple modifications, likewise introduced separately to all sequences, were employed to lock folds into one of the topologies without noticeable structural alterations. The unique combination of sequence mutations, high-resolution NMR structural information, and the thermodynamic stability for both topological competitors identified critical loop residue interactions. In contrast to first loop residues, which are mostly disordered and exposed to solvent in both propeller and lateral loops bridging a narrow groove, the last loop residue in a lateral three-nucleotide loop is engaged in stabilizing stacking interactions. The propensity of single-nucleotide loops to favor all-parallel topologies by enforcing a propeller-like conformation of an additional longer loop is shown to result from their preference in linking two outer tetrads of the same tetrad polarity. Taken together, the present studies contribute to a better structural and thermodynamic understanding of delicate loop interactions in genomic and artificially designed quadruplexes, e.g. when employed as therapeutics or in other biotechnological applications.
- Institute of Biochemistry, Universität Greifswald, D-17489 Greifswald, Germany.
Organizational Affiliation: