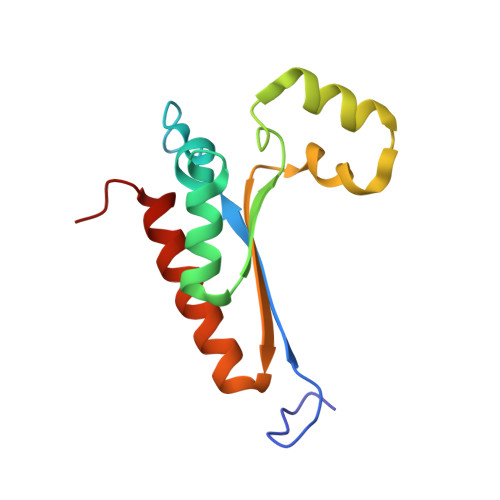
Structure and function analysis of Sam68 and hnRNP A1 synergy in the exclusion of exon 7 from SMN2 transcripts.
Nadal, M., Anton, R., Dorca-Arevalo, J., Estebanez-Perpina, E., Tizzano, E.F., Fuentes-Prior, P.(2023) Protein Sci 32: e4553-e4553
- PubMed: 36560896 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4553
- Primary Citation Related Structures:
7Z89, 7Z8A, 7Z9A, 7Z9B, 7ZAB, 7ZAC, 7ZAF, 7ZAM - PubMed Abstract:
Spinal muscular atrophy (SMA) is a neurodegenerative disease caused by the absence of a functional copy of the Survival of Motor Neuron 1 gene (SMN1). The nearly identical paralog, SMN2, cannot compensate for the loss of SMN1 because exon 7 is aberrantly skipped from most SMN2 transcripts, a process mediated by synergistic activities of Src-associated during mitosis, 68 kDa (Sam68/KHDRBS1) and heterogeneous nuclear ribonucleoprotein (hnRNP) A1. This results in the production of a truncated, nonfunctional protein that is rapidly degraded. Here, we present several crystal structures of Sam68 RNA-binding domain (RBD). Sam68-RBD forms stable symmetric homodimers by antiparallel association of helices α3 from two monomers. However, the details of domain organization and the dimerization interface differ significantly from previously characterized homologs. We demonstrate that Sam68 and hnRNP A1 can simultaneously bind proximal motifs within the central region of SMN2 (ex7). Furthermore, we show that the RNA-binding pockets of the two proteins are close to each other in their heterodimeric complex and identify contact residues using crosslinking-mass spectrometry. We present a model of the ternary Sam68·SMN2 (ex7)·hnRNP A1 complex that reconciles all available information on SMN1/2 splicing. Our findings have important implications for the etiology of SMA and open new avenues for the design of novel therapeutics to treat splicing diseases.
- Molecular Bases of Disease, Biomedical Research Institute Sant Pau (IIB Sant Pau), Barcelona, Spain.
Organizational Affiliation: