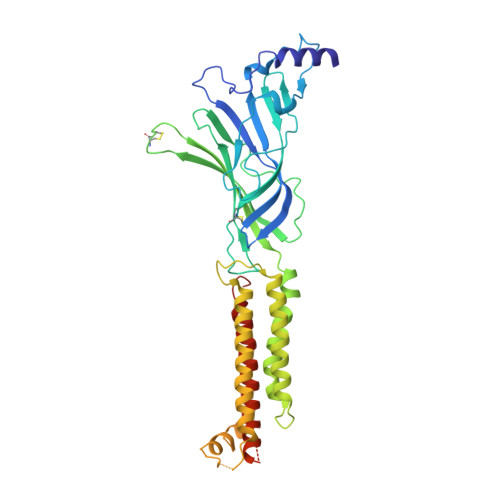
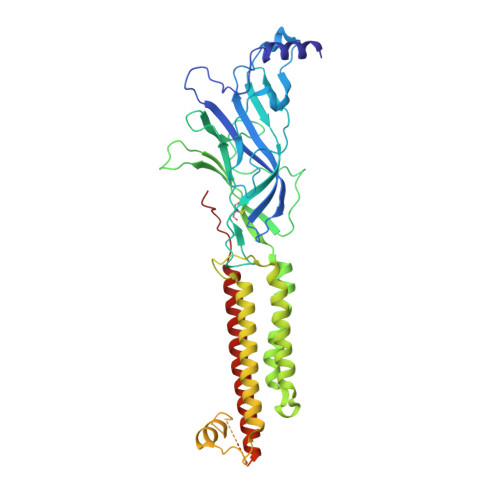
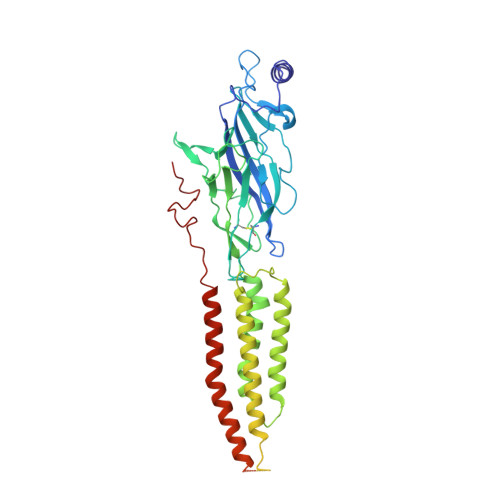
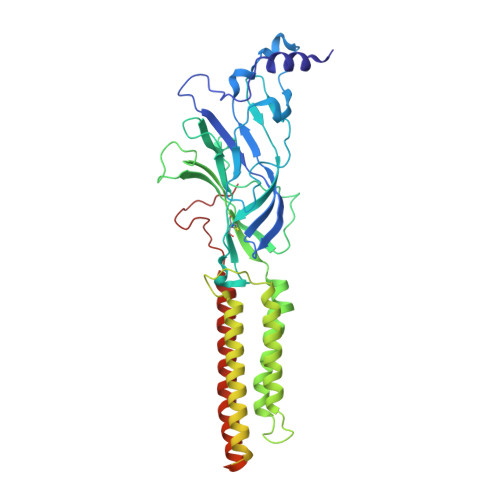
The molecular mechanism of snake short-chain alpha-neurotoxin binding to muscle-type nicotinic acetylcholine receptors.
Nys, M., Zarkadas, E., Brams, M., Mehregan, A., Kambara, K., Kool, J., Casewell, N.R., Bertrand, D., Baenziger, J.E., Nury, H., Ulens, C.(2022) Nat Commun 13: 4543-4543
- PubMed: 35927270 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-32174-7
- Primary Citation Related Structures:
7Z14 - PubMed Abstract:
Bites by elapid snakes (e.g. cobras) can result in life-threatening paralysis caused by venom neurotoxins blocking neuromuscular nicotinic acetylcholine receptors. Here, we determine the cryo-EM structure of the muscle-type Torpedo receptor in complex with ScNtx, a recombinant short-chain α-neurotoxin. ScNtx is pinched between loop C on the principal subunit and a unique hairpin in loop F on the complementary subunit, thereby blocking access to the neurotransmitter binding site. ScNtx adopts a binding mode that is tilted toward the complementary subunit, forming a wider network of interactions than those seen in the long-chain α-Bungarotoxin complex. Certain mutations in ScNtx at the toxin-receptor interface eliminate inhibition of neuronal α7 nAChRs, but not of human muscle-type receptors. These observations explain why ScNtx binds more tightly to muscle-type receptors than neuronal receptors. Together, these data offer a framework for understanding subtype-specific actions of short-chain α-neurotoxins and inspire strategies for design of new snake antivenoms.
- Laboratory of Structural Neurobiology, Department of Cellular and Molecular Medicine, Faculty of Medicine, KU Leuven, 3000, Leuven, Belgium. mieke.nys@kuleuven.be.
Organizational Affiliation: