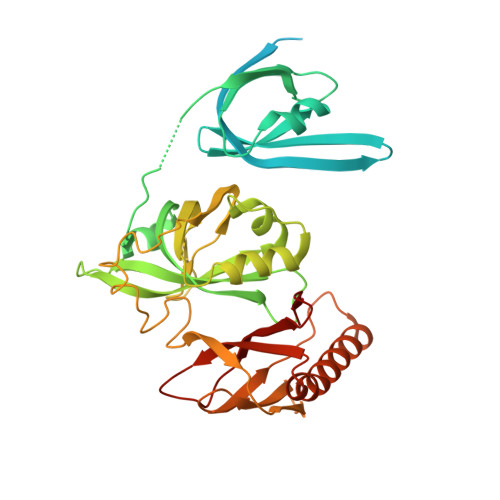
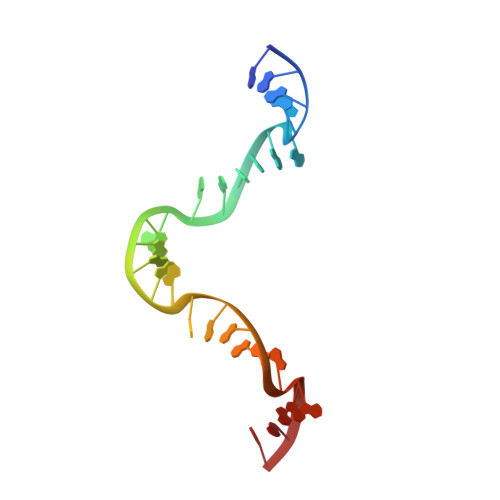
Structural transitions during the cooperative assembly of baculovirus single-stranded DNA-binding protein on ssDNA.
Yin, J., Fu, Y., Rao, G., Li, Z., Tian, K., Chong, T., Kuang, K., Wang, M., Hu, Z., Cao, S.(2022) Nucleic Acids Res 50: 13100-13113
- PubMed: 36477586 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac1142
- Primary Citation Related Structures:
7YNY, 7YPO, 7YPQ - PubMed Abstract:
Single-stranded DNA-binding proteins (SSBs) interact with single-stranded DNA (ssDNA) to form filamentous structures with various degrees of cooperativity, as a result of intermolecular interactions between neighboring SSB subunits on ssDNA. However, it is still challenging to perform structural studies on SSB-ssDNA filaments at high resolution using the most studied SSB models, largely due to the intrinsic flexibility of these nucleoprotein complexes. In this study, HaLEF-3, an SSB protein from Helicoverpa armigera nucleopolyhedrovirus, was used for in vitro assembly of SSB-ssDNA filaments, which were structurally studied at atomic resolution using cryo-electron microscopy. Combined with the crystal structure of ssDNA-free HaLEF-3 octamers, our results revealed that the three-dimensional rearrangement of HaLEF-3 induced by an internal hinge-bending movement is essential for the formation of helical SSB-ssDNA complexes, while the contacting interface between adjacent HaLEF-3 subunits remains basically intact. We proposed a local cooperative SSB-ssDNA binding model, in which, triggered by exposure to oligonucleotides, HaLEF-3 molecules undergo ring-to-helix transition to initiate continuous SSB-SSB interactions along ssDNA. Unique structural features revealed by the assembly of HaLEF-3 on ssDNA suggest that HaLEF-3 may represent a new class of SSB.
- CAS Key Laboratory of Special Pathogens, Wuhan Institute of Virology, Center for Biosafety Mega-Science, Chinese Academy of Sciences, Wuhan 430071, PR China.
Organizational Affiliation: