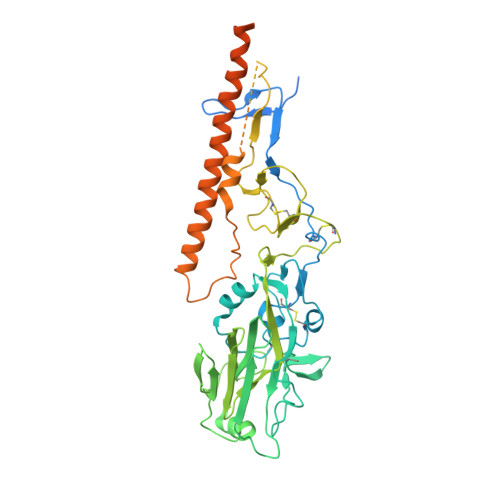
Identification of a cross-neutralizing antibody that targets the receptor binding site of H1N1 and H5N1 influenza viruses.
Li, T., Chen, J., Zheng, Q., Xue, W., Zhang, L., Rong, R., Zhang, S., Wang, Q., Hong, M., Zhang, Y., Cui, L., He, M., Lu, Z., Zhang, Z., Chi, X., Li, J., Huang, Y., Wang, H., Tang, J., Ying, D., Zhou, L., Wang, Y., Yu, H., Zhang, J., Gu, Y., Chen, Y., Li, S., Xia, N.(2022) Nat Commun 13: 5182-5182
- PubMed: 36056024 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-32926-5
- Primary Citation Related Structures:
7FAH, 7YHK - PubMed Abstract:
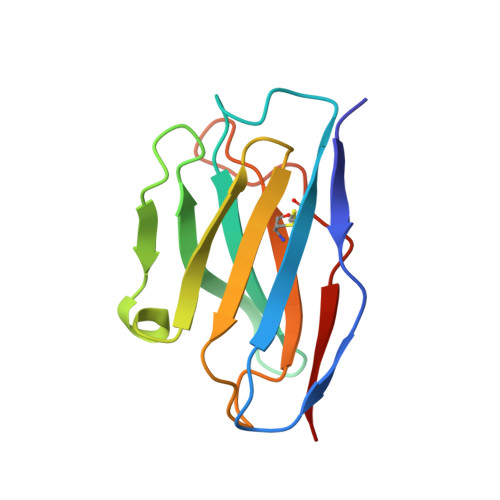
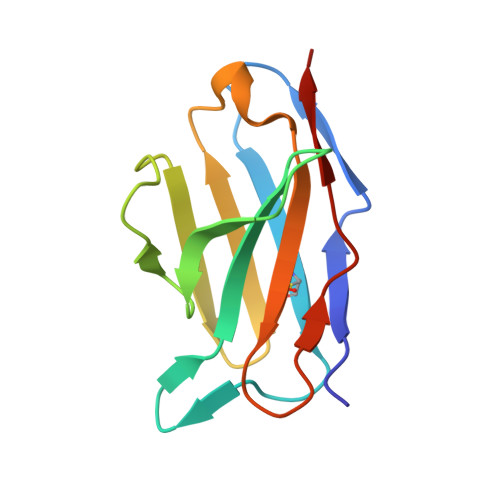
Influenza A viruses pose a significant threat globally each year, underscoring the need for a vaccine- or antiviral-based broad-protection strategy. Here, we describe a chimeric monoclonal antibody, C12H5, that offers neutralization against seasonal and pandemic H1N1 viruses, and cross-protection against some H5N1 viruses. Notably, C12H5 mAb offers broad neutralizing activity against H1N1 and H5N1 viruses by controlling virus entry and egress, and offers protection against H1N1 and H5N1 viral challenge in vivo. Through structural analyses, we show that C12H5 engages hemagglutinin (HA), the major surface glycoprotein on influenza, at a distinct epitope overlapping the receptor binding site and covering the 140-loop. We identified eight highly conserved (~90%) residues that are essential for broad H1N1 recognition, with evidence of tolerance for Asp or Glu at position 190; this site is a molecular determinant for human or avian host-specific recognition and this tolerance endows C12H5 with cross-neutralization potential. Our results could benefit the development of antiviral drugs and the design of broad-protection influenza vaccines.
- State Key Laboratory of Molecular Vaccinology and Molecular Diagnostics, School of Life Sciences, School of Public Health, Xiamen University, 361102, Xiamen, Fujian, China.
Organizational Affiliation: