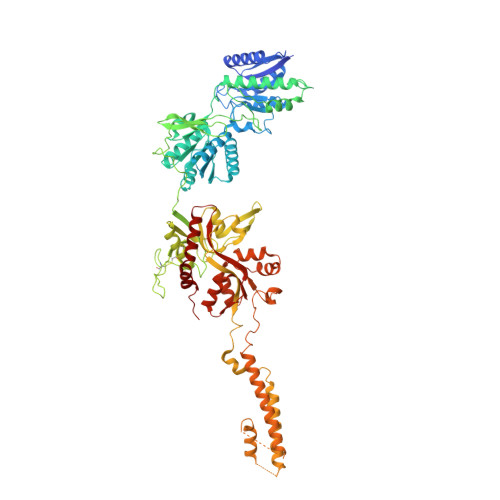
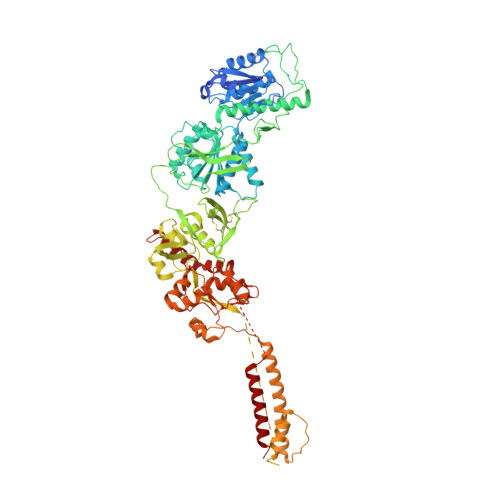
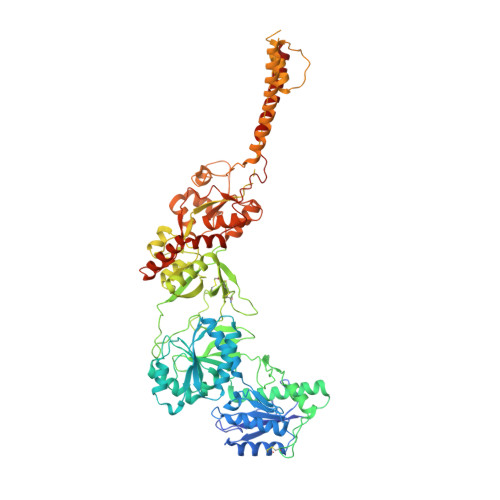
Distinct structure and gating mechanism in diverse NMDA receptors with GluN2C and GluN2D subunits.
Zhang, J., Zhang, M., Wang, Q., Wen, H., Liu, Z., Wang, F., Wang, Y., Yao, F., Song, N., Kou, Z., Li, Y., Guo, F., Zhu, S.(2023) Nat Struct Mol Biol 30: 629-639
- PubMed: 36959261 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-023-00959-z
- Primary Citation Related Structures:
7YFF, 7YFG, 7YFH, 7YFI, 7YFL, 7YFM, 7YFO, 7YFR, 8HDK - PubMed Abstract:
N-methyl-D-aspartate (NMDA) receptors are heterotetramers comprising two GluN1 and two alternate GluN2 (N2A-N2D) subunits. Here we report full-length cryo-EM structures of the human N1-N2D di-heterotetramer (di-receptor), rat N1-N2C di-receptor and N1-N2A-N2C tri-heterotetramer (tri-receptor) at a best resolution of 3.0 Å. The bilobate N-terminal domain (NTD) in N2D intrinsically adopts a closed conformation, leading to a compact NTD tetramer in the N1-N2D receptor. Additionally, crosslinking the ligand-binding domain (LBD) of two N1 protomers significantly elevated the channel open probability (Po) in N1-N2D di-receptors. Surprisingly, the N1-N2C di-receptor adopted both symmetric (minor) and asymmetric (major) conformations, the latter further locked by an allosteric potentiator, PYD-106, binding to a pocket between the NTD and LBD in only one N2C protomer. Finally, the N2A and N2C subunits in the N1-N2A-N2C tri-receptor display a conformation close to one protomer in the N1-N2A and N1-N2C di-receptors, respectively. These findings provide a comprehensive structural understanding of diverse function in major NMDA receptor subtypes.
- Institute of Neuroscience, State Key Laboratory of Neuroscience, CAS Center for Excellence in Brain Science and Intelligence Technology, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: