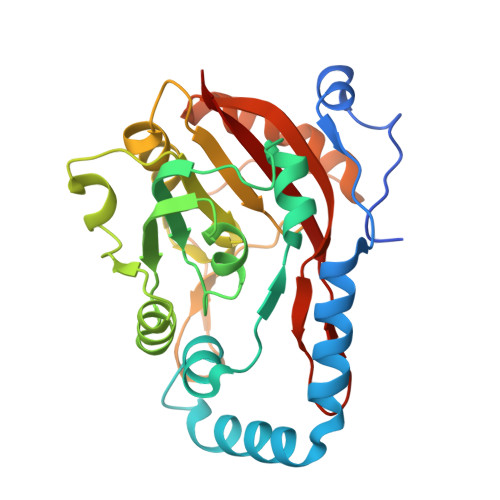
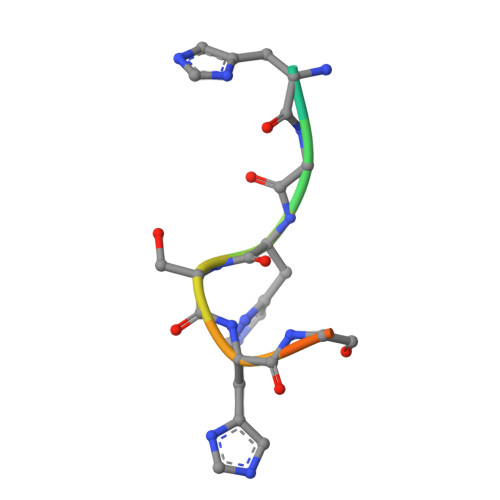
Molecular basis for METTL9-mediated N1-histidine methylation.
Wang, X., Xie, H., Guo, Q., Cao, D., Ru, W., Zhao, S., Zhu, Z., Zhang, J., Pan, W., Yao, X., Xu, C.(2023) Cell Discov 9: 38-38
- PubMed: 37015930 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-023-00548-w
- Primary Citation Related Structures:
7Y9C, 7YF2, 7YF3, 7YF4 - MOE Key Laboratory for Membraneless Organelles and Cellular Dynamics, Hefei National Research Center for Interdisciplinary Sciences at the Microscale, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, China.
Organizational Affiliation: