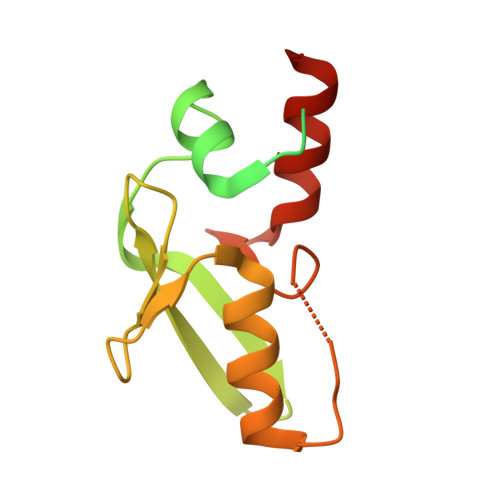
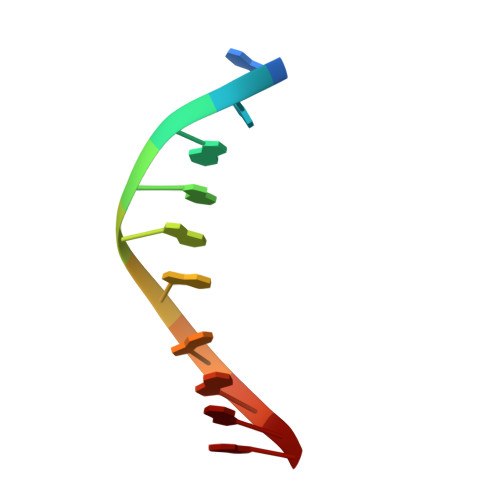
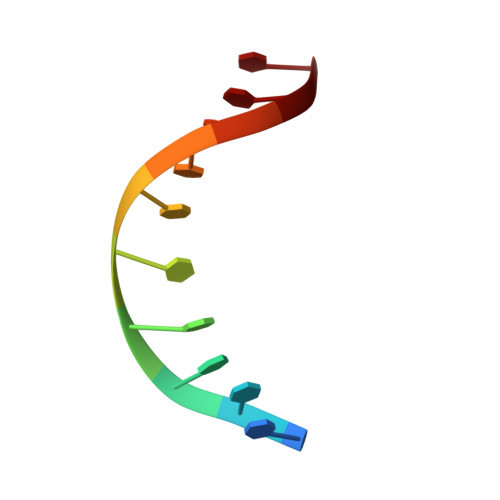
Structures of MPND Reveal the Molecular Recognition of Nucleosomes.
Yang, M., Li, X., Tian, Z., Ma, L., Ma, J., Liu, Y., Shang, G., Liang, A., Wu, W., Chen, Z.(2023) Int J Mol Sci 24
- PubMed: 36834777 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms24043368
- Primary Citation Related Structures:
7YDT, 7YDW - PubMed Abstract:
Adenine N 6 methylation in DNA (6mA) is a well-known epigenetic modification in bacteria, phages, and eukaryotes. Recent research has identified the Mpr1/Pad1 N-terminal (MPN) domain-containing protein (MPND) as a sensor protein that may recognize DNA 6mA modification in eukaryotes. However, the structural details of MPND and the molecular mechanism of their interaction remain unknown. Herein, we report the first crystal structures of the apo-MPND and MPND-DNA complex at resolutions of 2.06 Å and 2.47 Å, respectively. In solution, the assemblies of both apo-MPND and MPND-DNA are dynamic. In addition, MPND was found to possess the ability to bind directly to histones, no matter the N-terminal restriction enzyme-adenine methylase-associated domain or the C-terminal MPN domain. Moreover, the DNA and the two acidic regions of MPND synergistically enhance the interaction between MPND and histones. Therefore, our findings provide the first structural information regarding the MPND-DNA complex and also provide evidence of MPND-nucleosome interactions, thereby laying the foundation for further studies on gene control and transcriptional regulation.
- State Key Laboratory of Agrobiotechnology, College of Biological Sciences, China Agricultural University, Beijing 100193, China.
Organizational Affiliation: