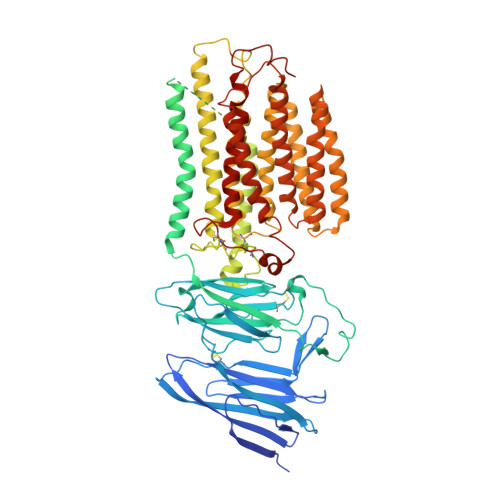
Structural insight into the human SID1 transmembrane family member 2 reveals its lipid hydrolytic activity.
Qian, D., Cong, Y., Wang, R., Chen, Q., Yan, C., Gong, D.S.(2023) Nat Commun 14: 3568-3568
- PubMed: 37322007 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-39335-2
- Primary Citation Related Structures:
7Y63, 7Y68, 7Y69 - PubMed Abstract:
The systemic RNAi-defective (SID) transmembrane family member 2 (SIDT2) is a putative nucleic acid channel or transporter that plays essential roles in nucleic acid transport and lipid metabolism. Here, we report the cryo-electron microscopy (EM) structures of human SIDT2, which forms a tightly packed dimer with extensive interactions mediated by two previously uncharacterized extracellular/luminal β-strand-rich domains and the unique transmembrane domain (TMD). The TMD of each SIDT2 protomer contains eleven transmembrane helices (TMs), and no discernible nucleic acid conduction pathway has been identified within the TMD, suggesting that it may act as a transporter. Intriguingly, TM3-6 and TM9-11 form a large cavity with a putative catalytic zinc atom coordinated by three conserved histidine residues and one aspartate residue lying approximately 6 Å from the extracellular/luminal surface of the membrane. Notably, SIDT2 can hydrolyze C18 ceramide into sphingosine and fatty acid with a slow rate. The information presented advances the understanding of the structure-function relationships in the SID1 family proteins.
- State Key Laboratory of Medicinal Chemical Biology and College of Life Sciences, Nankai University, Tianjin, 300350, China.
Organizational Affiliation: