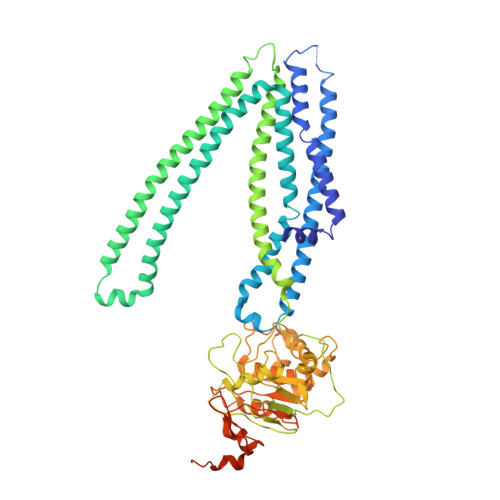
Cryo-EM structures of mitochondrial ABC transporter ABCB10 in apo and biliverdin-bound form.
Cao, S., Yang, Y., He, L., Hang, Y., Yan, X., Shi, H., Wu, J., Ouyang, Z.(2023) Nat Commun 14: 2030-2030
- PubMed: 37041204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-37851-9
- Primary Citation Related Structures:
7Y48, 7Y49 - PubMed Abstract:
ABCB10, a member of ABC transporter superfamily that locates in the inner membrane of mitochondria, plays crucial roles in hemoglobin synthesis, antioxidative stress and stabilization of the iron transporter mitoferrin-1. Recently, it was found that ABCB10 is a mitochondrial biliverdin exporter. However, the molecular mechanism of biliverdin export by ABCB10 remains elusive. Here we report the cryo-EM structures of ABCB10 in apo (ABCB10-apo) and biliverdin-bound form (ABCB10-BV) at 3.67 Å and 2.85 Å resolution, respectively. ABCB10-apo adopts a wide-open conformation and may thus represent the apo form structure. ABCB10-BV forms a closed conformation and biliverdin situates in a hydrophobic pocket in one protomer and bridges the interaction through hydrogen bonds with the opposing one. We also identify cholesterols sandwiched by BVs and discuss the export dynamics based on these structural and biochemical observations.
- Wuxi Biortus Biosciences Co. Ltd., 6 Dongsheng Western Road, 214437, Jiangyin, Jiangsu, China.
Organizational Affiliation: