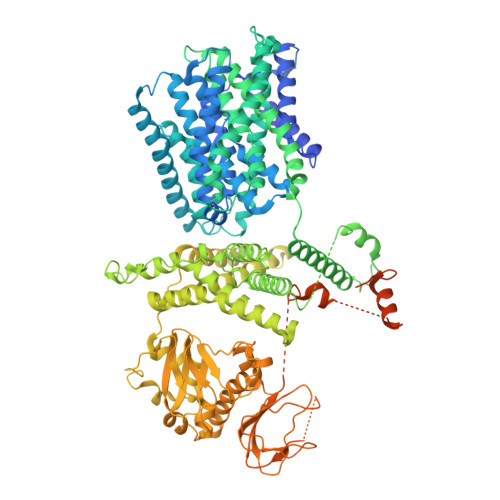
Architecture and autoinhibitory mechanism of the plasma membrane Na + /H + antiporter SOS1 in Arabidopsis.
Wang, Y., Pan, C., Chen, Q., Xie, Q., Gao, Y., He, L., Li, Y., Dong, Y., Jiang, X., Zhao, Y.(2023) Nat Commun 14: 4487-4487
- PubMed: 37495621 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-40215-y
- Primary Citation Related Structures:
7Y3E, 8HYA - PubMed Abstract:
Salt-overly-sensitive 1 (SOS1) is a unique electroneutral Na + /H + antiporter at the plasma membrane of higher plants and plays a central role in resisting salt stress. SOS1 is kept in a resting state with basal activity and activated upon phosphorylation. Here, we report the structures of SOS1. SOS1 forms a homodimer, with each monomer composed of transmembrane and intracellular domains. We find that SOS1 is locked in an occluded state by shifting of the lateral-gate TM5b toward the dimerization domain, thus shielding the Na + /H + binding site. We speculate that the dimerization of the intracellular domain is crucial to stabilize the transporter in this specific conformation. Moreover, two discrete fragments and a residue W1013 are important to prevent the transition of SOS1 to an alternative conformational state, as validated by functional complementation assays. Our study enriches understanding of the alternate access model of eukaryotic Na + /H + exchangers.
- National Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 100101, Beijing, China.
Organizational Affiliation: