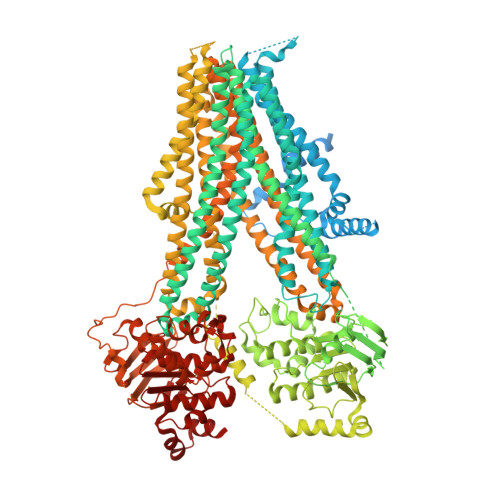
The inhibition mechanism of the SUR2A-containing K ATP channel by a regulatory helix.
Ding, D., Hou, T., Wei, M., Wu, J.X., Chen, L.(2023) Nat Commun 14: 3608-3608
- PubMed: 37330603
- DOI: https://doi.org/10.1038/s41467-023-39379-4
- Primary Citation of Related Structures:
7Y1J, 7Y1K, 7Y1L, 7Y1M, 7Y1N - PubMed Abstract:
K ATP channels are metabolic sensors for intracellular ATP/ADP ratios, play essential roles in many physiological processes, and are implicated in a spectrum of pathological conditions. SUR2A-containing K ATP channels differ from other subtypes in their sensitivity to Mg-ADP activation. However, the underlying structural mechanism remains poorly understood. Here we present a series of cryo-EM structures of SUR2A in the presence of different combinations of Mg-nucleotides and the allosteric inhibitor repaglinide. These structures uncover regulatory helix (R helix) on the NBD1-TMD2 linker, which wedges between NBD1 and NBD2. R helix stabilizes SUR2A in the NBD-separated conformation to inhibit channel activation. The competitive binding of Mg-ADP with Mg-ATP to NBD2 mobilizes the R helix to relieve such inhibition, allowing channel activation. The structures of SUR2B in similar conditions suggest that the C-terminal 42 residues of SUR2B enhance the structural dynamics of NBD2 and facilitate the dissociation of the R helix and the binding of Mg-ADP to NBD2, promoting NBD dimerization and subsequent channel activation.
- State Key Laboratory of Membrane Biology, College of Future Technology, Institute of Molecular Medicine, Peking University, Beijing Key Laboratory of Cardiometabolic Molecular Medicine, 100871, Beijing, China.
Organizational Affiliation: