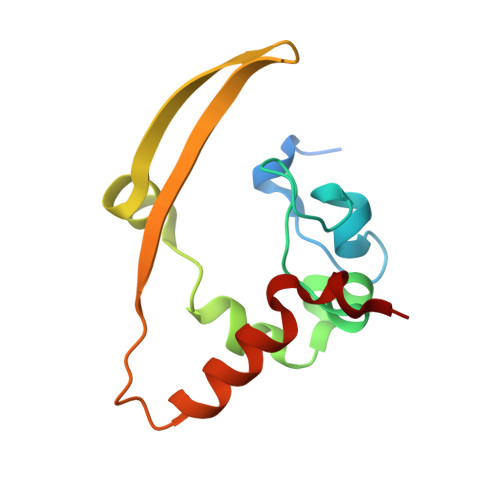
Structural basis of the C-terminal domain of SARS-CoV-2 N protein in complex with GMP reveals critical residues for RNA interaction.
Ni, X., Han, Y., Yu, J., Zhou, R., Lei, J.(2024) Bioorg Med Chem Lett : 130014-130014
- PubMed: 39489230 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2024.130014
- Primary Citation Related Structures:
7XXK - PubMed Abstract:
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) nucleocapsid (N) protein performs multiple functions during the viral life cycle, particularly in binding to the viral genomic RNA to form a helical ribonucleoprotein complex. Here, we present that the C-terminal domain of SARS-CoV-2 N protein (N-CTD) specifically interacts with polyguanylic acid (poly(G)). The crystal structure of the N-CTD in complex with 5'-guanylic acid (GMP, also known as guanosine monophosphate) was determined at a resolution of approximately 2.0 Å. A novel GMP-binding pocket in the N-CTD was illustrated. Residues Arg259 and Lys338 were identified to play key roles in binding to GMP through mutational analysis. These two residues are absolutely conserved in the other two highly pathogenic CoVs, SARS-CoV and Middle East respiratory syndrome coronavirus (MERS-CoV). Overall, our findings expand the structural information on N protein interacting with guanylate and reveal a conserved GMP-binding pocket as a potential antiviral target.
- National Clinical Research Center for Geriatrics, and State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, Sichuan, China.
Organizational Affiliation: