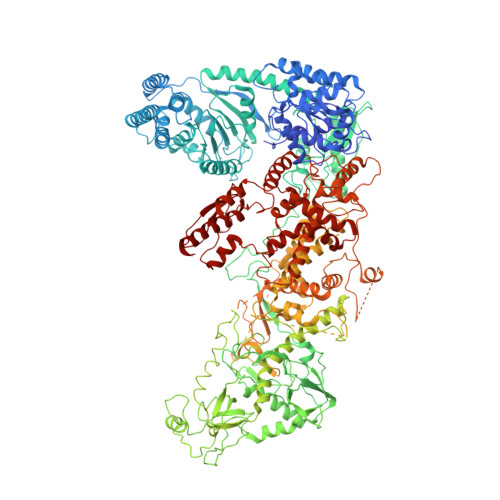
Structure of the human DICER-pre-miRNA complex in a dicing state.
Lee, Y.Y., Lee, H., Kim, H., Kim, V.N., Roh, S.H.(2023) Nature 615: 331-338
- PubMed: 36813958
- DOI: https://doi.org/10.1038/s41586-023-05723-3
- Primary Citation Related Structures:
7XW2, 7XW3 - PubMed Abstract:
Dicer has a key role in small RNA biogenesis, processing double-stranded RNAs (dsRNAs) 1,2 . Human DICER (hDICER, also known as DICER1) is specialized for cleaving small hairpin structures such as precursor microRNAs (pre-miRNAs) and has limited activity towards long dsRNAs-unlike its homologues in lower eukaryotes and plants, which cleave long dsRNAs. Although the mechanism by which long dsRNAs are cleaved has been well documented, our understanding of pre-miRNA processing is incomplete because structures of hDICER in a catalytic state are lacking. Here we report the cryo-electron microscopy structure of hDICER bound to pre-miRNA in a dicing state and uncover the structural basis of pre-miRNA processing. hDICER undergoes large conformational changes to attain the active state. The helicase domain becomes flexible, which allows the binding of pre-miRNA to the catalytic valley. The double-stranded RNA-binding domain relocates and anchors pre-miRNA in a specific position through both sequence-independent and sequence-specific recognition of the newly identified 'GYM motif' 3 . The DICER-specific PAZ helix is also reoriented to accommodate the RNA. Furthermore, our structure identifies a configuration of the 5' end of pre-miRNA inserted into a basic pocket. In this pocket, a group of arginine residues recognize the 5' terminal base (disfavouring guanine) and terminal monophosphate; this explains the specificity of hDICER and how it determines the cleavage site. We identify cancer-associated mutations in the 5' pocket residues that impair miRNA biogenesis. Our study reveals how hDICER recognizes pre-miRNAs with stringent specificity and enables a mechanistic understanding of hDICER-related diseases.
- Center for RNA Research, Institute for Basic Science (IBS), Seoul, Republic of Korea.
Organizational Affiliation: