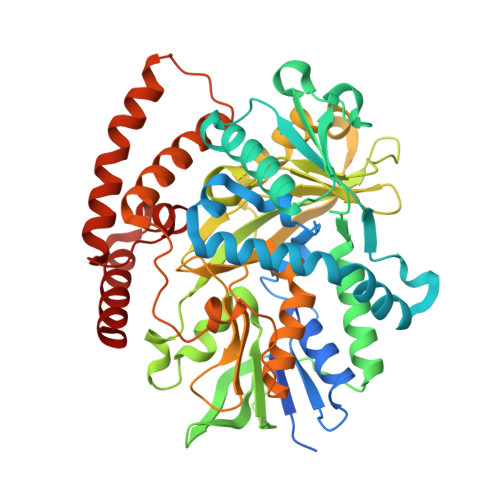
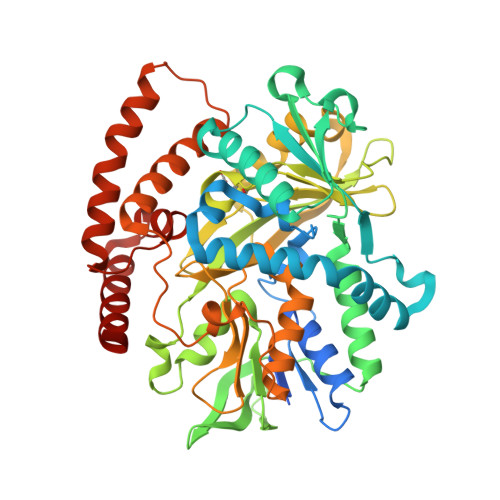
N-Formimidoylation/-iminoacetylation modification in aminoglycosides requires FAD-dependent and ligand-protein NOS bridge dual chemistry.
Wang, Y.L., Chang, C.Y., Hsu, N.S., Lo, I.W., Lin, K.H., Chen, C.L., Chang, C.F., Wang, Z.C., Ogasawara, Y., Dairi, T., Maruyama, C., Hamano, Y., Li, T.L.(2023) Nat Commun 14: 2528-2528
- PubMed: 37137912 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-38218-w
- Primary Citation Related Structures:
7XQA, 7XX0, 7XXC, 7XXD, 7XXM, 7XXP, 7XXR, 7XYE, 7XYL, 7Y0X, 7YPU, 7YPV, 8GRI - PubMed Abstract:
Oxidized cysteine residues are highly reactive and can form functional covalent conjugates, of which the allosteric redox switch formed by the lysine-cysteine NOS bridge is an example. Here, we report a noncanonical FAD-dependent enzyme Orf1 that adds a glycine-derived N-formimidoyl group to glycinothricin to form the antibiotic BD-12. X-ray crystallography was used to investigate this complex enzymatic process, which showed Orf1 has two substrate-binding sites that sit 13.5 Å apart unlike canonical FAD-dependent oxidoreductases. One site could accommodate glycine and the other glycinothricin or glycylthricin. Moreover, an intermediate-enzyme adduct with a NOS-covalent linkage was observed in the later site, where it acts as a two-scissile-bond linkage facilitating nucleophilic addition and cofactor-free decarboxylation. The chain length of nucleophilic acceptors vies with bond cleavage sites at either N-O or O-S accounting for N-formimidoylation or N-iminoacetylation. The resultant product is no longer sensitive to aminoglycoside-modifying enzymes, a strategy that antibiotic-producing species employ to counter drug resistance in competing species.
- Genomics Research Center, Academia Sinica, Taipei, 11529, Taiwan.
Organizational Affiliation: