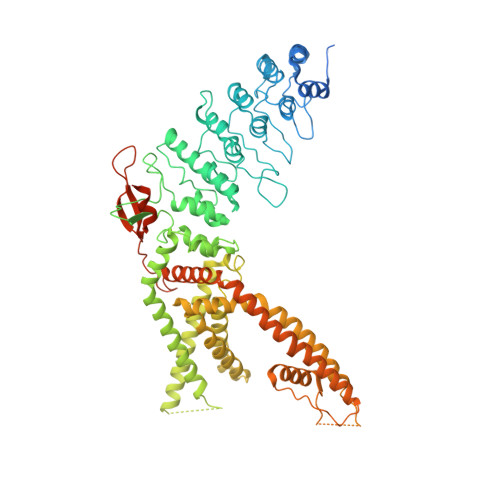
Structural mechanisms of TRPV2 modulation by endogenous and exogenous ligands.
Su, N., Zhen, W., Zhang, H., Xu, L., Jin, Y., Chen, X., Zhao, C., Wang, Q., Wang, X., Li, S., Wen, H., Yang, W., Guo, J., Yang, F.(2023) Nat Chem Biol 19: 72-80
- PubMed: 36163384 Search on PubMed
- DOI: https://doi.org/10.1038/s41589-022-01139-8
- Primary Citation Related Structures:
7XEM, 7XEO, 7XER, 7XEU, 7XEV, 7XEW, 7YEP - PubMed Abstract:
The transient receptor potential vanilloid 2 (TRPV2) ion channel is a polymodal receptor widely involved in many physiological and pathological processes. Despite many TRPV2 modulators being identified, whether and how TRPV2 is regulated by endogenous lipids remains elusive. Here, we report an endogenous cholesterol molecule inside the vanilloid binding pocket (VBP) of TRPV2, with a 'head down, tail up' configuration, resolved at 3.2 Å using cryo-EM. Cholesterol binding antagonizes ligand activation of TRPV2, which is removed from VBP by methyl-β-cyclodextrin (MβCD) as resolved at 2.9 Å. We also observed that estradiol (E2) potentiated TRPV2 activation by 2-aminoethoxydiphenyl borate (2-APB), a classic tool compound for TRP channels. Our cryo-EM structures (resolved at 2.8-3.3 Å) further suggest how E2 disturbed cholesterol binding and how 2-APB bound within the VBP with E2 or without both E2 and endogenous cholesterol, respectively. Therefore, our study has established the structural basis for ligand recognition of the inhibitory endogenous cholesterol and excitatory exogenous 2-APB in TRPV2.
- Department of Biophysics, and Kidney Disease Center of the First Affiliated Hospital, Zhejiang University School of Medicine, Hangzhou, China.
Organizational Affiliation: