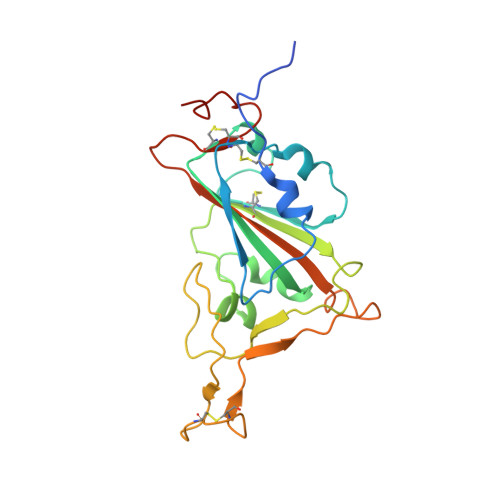
Host range and structural analysis of bat-origin RshSTT182/200 coronavirus binding to human ACE2 and its animal orthologs.
Hu, Y., Liu, K., Han, P., Xu, Z., Zheng, A., Pan, X., Jia, Y., Su, C., Tang, L., Wu, L., Bai, B., Zhao, X., Tian, D., Chen, Z., Qi, J., Wang, Q., Gao, G.F.(2023) EMBO J 42: e111737-e111737
- PubMed: 36519268 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2022111737
- Primary Citation Related Structures:
7XBF, 7XBG, 7XBH - PubMed Abstract:
Bat-origin RshSTT182 and RshSTT200 coronaviruses (CoV) from Rhinolophus shameli in Southeast Asia (Cambodia) share 92.6% whole-genome identity with SARS-CoV-2 and show identical receptor-binding domains (RBDs). In this study, we determined the structure of the RshSTT182/200 receptor binding domain (RBD) in complex with human angiotensin-converting enzyme 2 (hACE2) and identified the key residues that influence receptor binding. The binding of the RshSTT182/200 RBD to ACE2 orthologs from 39 animal species, including 18 bat species, was used to evaluate its host range. The RshSTT182/200 RBD broadly recognized 21 of 39 ACE2 orthologs, although its binding affinities for the orthologs were weaker than those of the RBD of SARS-CoV-2. Furthermore, RshSTT182 pseudovirus could utilize human, fox, and Rhinolophus affinis ACE2 receptors for cell entry. Moreover, we found that SARS-CoV-2 induces cross-neutralizing antibodies against RshSTT182 pseudovirus. Taken together, these findings indicate that RshSTT182/200 can potentially infect susceptible animals, but requires further evolution to obtain strong interspecies transmission abilities like SARS-CoV-2.
- School of Life Sciences, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, China.
Organizational Affiliation: