Structural insights into the ligand binding and G i coupling of serotonin receptor 5-HT 5A .
Tan, Y., Xu, P., Huang, S., Yang, G., Zhou, F., He, X., Ma, H., Xu, H.E., Jiang, Y.(2022) Cell Discov 8: 50-50
- PubMed: 35610220 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-022-00412-3
- Primary Citation Related Structures:
7X5H - PubMed Abstract:
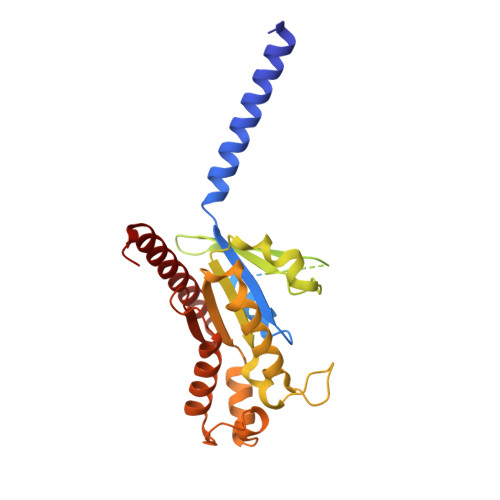
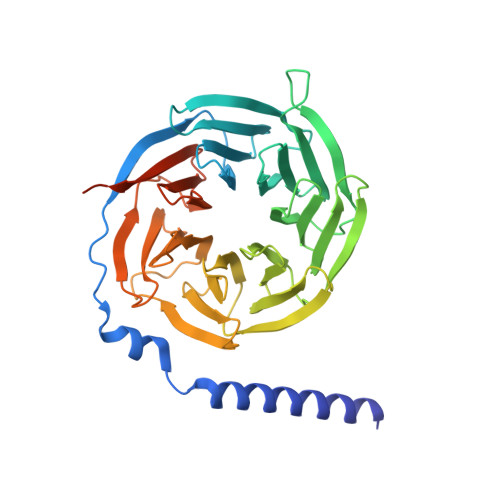
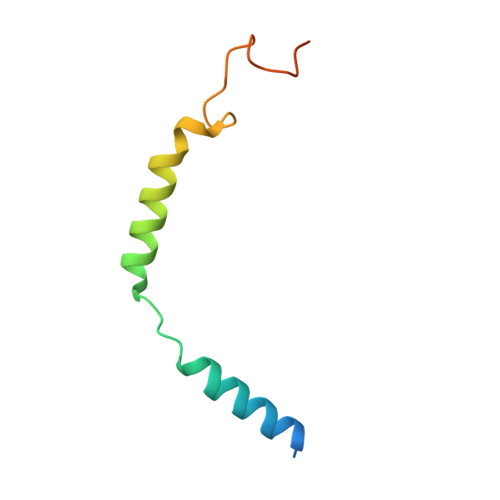
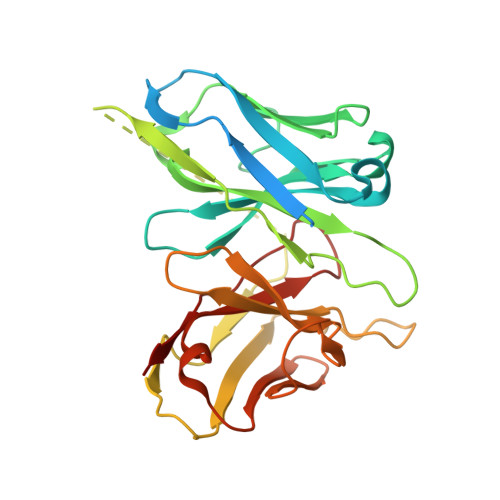
5-hydroxytryptamine receptor 5A (5-HT 5A ) belongs to the 5-HT receptor family and signals through the G i/o protein. It is involved in nervous system regulation and an attractive target for the treatment of psychosis, depression, schizophrenia, and neuropathic pain. 5-HT 5A is the only G i/o -coupled 5-HT receptor subtype lacking a high-resolution structure, which hampers the mechanistic understanding of ligand binding and G i/o coupling for 5-HT 5A . Here we report a cryo-electron microscopy structure of the 5-HT 5A -G i complex bound to 5-Carboxamidotryptamine (5-CT). Combined with functional analysis, this structure reveals the 5-CT recognition mechanism and identifies the receptor residue at 6.55 as a determinant of the 5-CT selectivity for G i/o -coupled 5-HT receptors. In addition, 5-HT 5A shows an overall conserved G i protein coupling mode compared with other G i/o -coupled 5-HT receptors. These findings provide comprehensive insights into the ligand binding and G protein coupling of G i/o -coupled 5-HT receptors and offer a template for the design of 5-HT 5A -selective drugs.
- The CAS Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: