Molecular basis of PD-1 blockade by dostarlimab, the FDA-approved antibody for cancer immunotherapy.
Park, U.B., Jeong, T.J., Gu, N., Lee, H.T., Heo, Y.S.(2022) Biochem Biophys Res Commun 599: 31-37
- PubMed: 35168061 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2022.02.026
- Primary Citation Related Structures:
7WSL - PubMed Abstract:
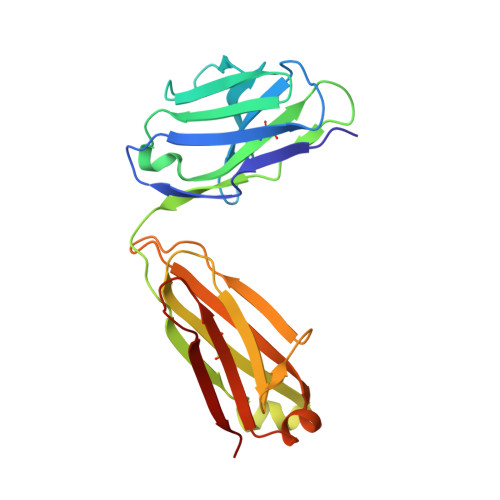
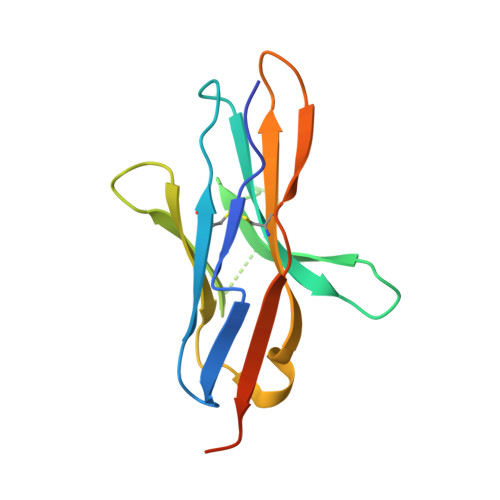
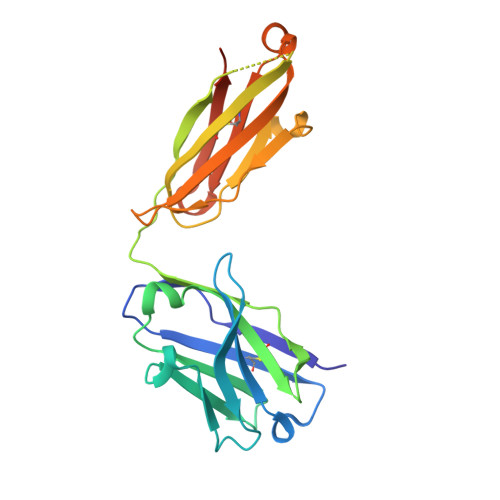
Targeting of programmed cell death 1 (PD-1) with monoclonal antibodies to block the interaction with its ligand PD-L1 has been successful in immunotherapy of multiple types of cancer, and their mechanism involves the restoration of the T-cell immune response. April 2021, the US FDA approved dostarlimab, a therapeutic antibody against PD-1, for the treatment of endometrial cancer. Here, we report the crystal structure of the extracellular domain of PD-1 in complex with the dostarlimab Fab at the resolution of 1.53 Å. Although the interaction between PD-1 and dostarlimab involves mainly the residues within the heavy chain of dostarlimab, the steric occlusion of PD-L1 binding is primarily contributed by the light chain. Dostarlimab induces conformational rearrangements of the BC, C'D and FG loops of PD-1 to achieve a high affinity. Significantly, the residue R86 within the C'D loop of PD-1 plays a critical role for dostarlimab binding by occupying the concave surface on the heavy chain via multiple interactions. This high-resolution structure can provide helpful information for designing improved anti-PD-1 biologics or effective combination strategies for cancer immunotherapy.
- Department of Chemistry, Konkuk University, 120 Neungdong-ro, Gwangjin-gu, Seoul, 05029, Republic of Korea.
Organizational Affiliation: