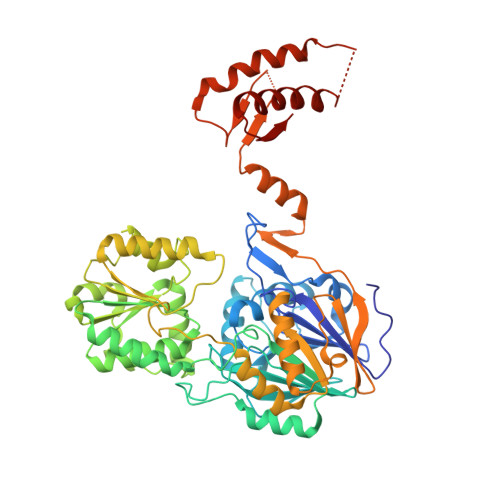
Structural insights into RNase J that plays an essential role in Mycobacterium tuberculosis RNA metabolism
Bao, L., Hu, J., Zhan, B., Chi, M., Li, Z., Wang, S., Shan, C., Zhao, Z., Guo, Y., Ding, X., Ji, C., Tao, S., Ni, T., Zhang, X., Zhao, G., Li, J.(2023) Nat Commun 14: 2280