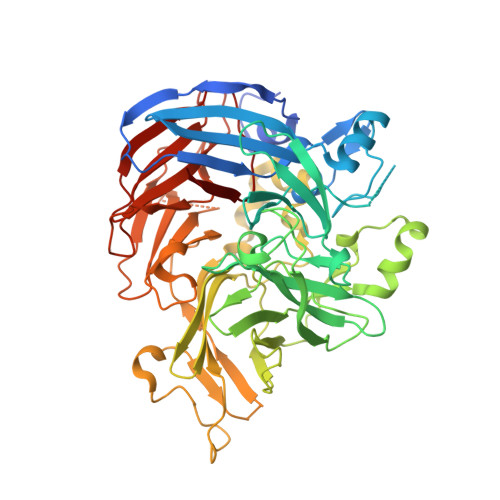
Structural and Functional Analysis of Nonheme Iron Enzymes BCMO-1 and BCMO-2 from Caenorhabditis elegans .
Pan, W., Zhou, Y.L., Wang, J., Dai, H.E., Wang, X., Liu, L.(2022) Front Mol Biosci 9: 844453-844453
- PubMed: 35223999 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fmolb.2022.844453
- Primary Citation Related Structures:
7WH0, 7WH1 - PubMed Abstract:
Carotenoid metabolism is critical for diverse physiological processes. The nematode Caenorhabditis elegans has two genes that are annotated as β-carotene 15,15'-monooxygenase (BCMO) and are 17 centimorgan apart on chromosome II, but the function of BCMO-1 and BCMO-2 remains uncharacterized. Sequence homology indicates that the two enzymes belong to the carotenoid cleavage dioxygenase family that share a seven-bladed β-propeller fold with a nonheme iron center. Here we determined crystal structures of BCMO-1 and BCMO-2 at resolutions of 1.8 and 1.9 Å, respectively. Structural analysis reveals that BCMO-1 and BCMO-2 are strikingly similar to each other. We also characterized their β-carotene cleavage activity, but the results suggest that they may not act as β-carotene 15,15'-oxygenases.
- School of Life Sciences, Anhui University, Hefei, China.
Organizational Affiliation: