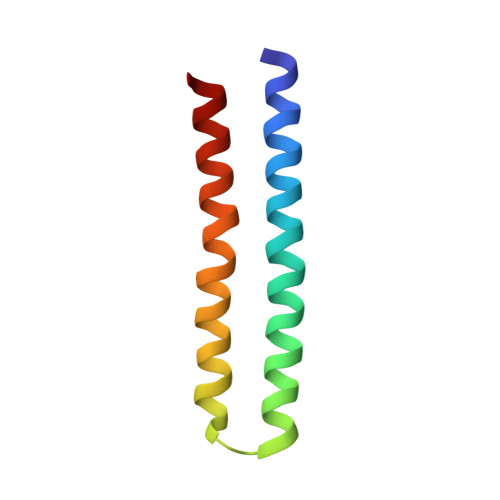
Chemical Conformation of the Essential Glutamate Site of the c -Ring within Thermophilic Bacillus F o F 1 -ATP Synthase Determined by Solid-State NMR Based on its Isolated c -Ring Structure.
Todokoro, Y., Kang, S.J., Suzuki, T., Ikegami, T., Kainosho, M., Yoshida, M., Fujiwara, T., Akutsu, H.(2022) J Am Chem Soc 144: 14132-14139
- PubMed: 35905443 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.2c03580
- Primary Citation Related Structures:
7WEM - PubMed Abstract:
Proton translocation through the membrane-embedded F o component of F-type ATP synthase (F o F 1 ) is facilitated by the rotation of the F o c -subunit ring ( c -ring), carrying protons at essential acidic amino acid residues. Cryo-electron microscopy (Cryo-EM) structures of F o F 1 suggest a unique proton translocation mechanism. To elucidate it based on the chemical conformation of the essential acidic residues of the c -ring in F o F 1 , we determined the structure of the isolated thermophilic Bacillus F o (tF o ) c -ring, consisting of 10 subunits, in membranes by solid-state NMR. This structure contains a distinct proton-locking conformation, wherein Asn23 ( c N23) C γ O and Glu56 ( c E56) C δ OH form a hydrogen bond in a closed form. We introduced stereo-array-isotope-labeled (SAIL) Glu and Asn into the tF o c -ring to clarify the chemical conformation of these residues in tF o F 1 -ATP synthase (tF o F 1 ). Two well-separated 13 C signals could be detected for c N23 and c E56 in a 505 kDa membrane protein complex, respectively, thereby suggesting the presence of two distinct chemical conformations. Based on the signal intensity and structure of the tF o c -ring and tF o F 1 , six pairs of c N23 and c E56 surrounded by membrane lipids take the closed form, whereas the other four in the a - c interface employ the deprotonated open form at a proportion of 87%. This indicates that the a - c interface is highly hydrophilic. The p K a values of the four c E56 residues in the a - c interface were estimated from the c N23 signal intensity in the open and closed forms and distribution of polar residues around each c E56. The results favor a rotation of the c -ring for ATP synthesis.
- Institute for Protein Research, Osaka University, 3-2 Yamadaoka, Suita 565-0871, Japan.
Organizational Affiliation: