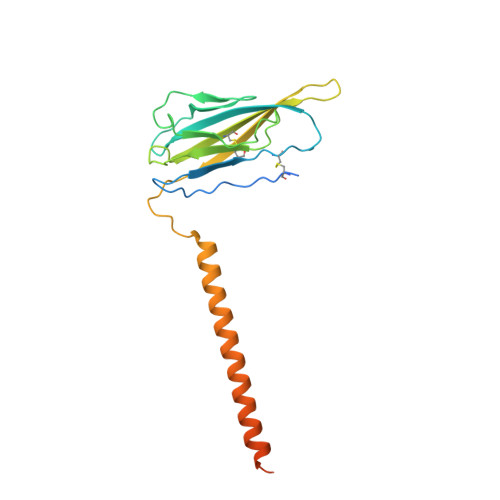
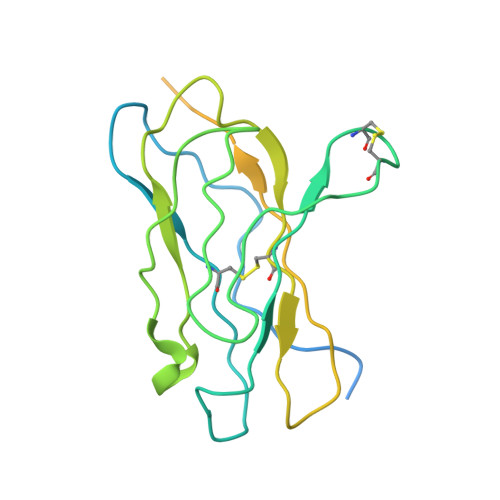
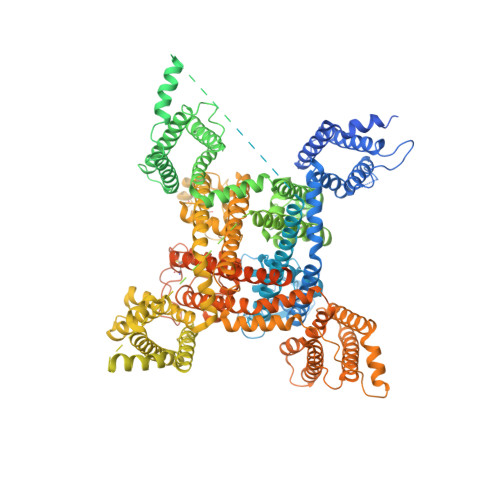
Structural basis for modulation of human Na V 1.3 by clinical drug and selective antagonist.
Li, X., Xu, F., Xu, H., Zhang, S., Gao, Y., Zhang, H., Dong, Y., Zheng, Y., Yang, B., Sun, J., Zhang, X.C., Zhao, Y., Jiang, D.(2022) Nat Commun 13: 1286-1286
- PubMed: 35277491 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-28808-5
- Primary Citation Related Structures:
7W77, 7W7F - PubMed Abstract:
Voltage-gated sodium (Na V ) channels play fundamental roles in initiating and propagating action potentials. Na V 1.3 is involved in numerous physiological processes including neuronal development, hormone secretion and pain perception. Here we report structures of human Na V 1.3/β1/β2 in complex with clinically-used drug bulleyaconitine A and selective antagonist ICA121431. Bulleyaconitine A is located around domain I-II fenestration, providing the detailed view of the site-2 neurotoxin binding site. It partially blocks ion path and expands the pore-lining helices, elucidating how the bulleyaconitine A reduces peak amplitude but improves channel open probability. In contrast, ICA121431 preferentially binds to activated domain IV voltage-sensor, consequently strengthens the Ile-Phe-Met motif binding to its receptor site, stabilizes the channel in inactivated state, revealing an allosterically inhibitory mechanism of Na V channels. Our results provide structural details of distinct small-molecular modulators binding sites, elucidate molecular mechanisms of their action on Na V channels and pave a way for subtype-selective therapeutic development.
- Laboratory of Soft Matter Physics, Institute of Physics, Chinese Academy of Sciences, Beijing, 100190, China.
Organizational Affiliation: